Instituto de Medicina Tropical "Pedro Kourí"
Dra. María T. Illnait Zaragozí,1 Lic. Yoel Miranda Toirac,2 Lic. Iliana del C. Valdés Hernández,3 Dr. Gerardo Martínez Machín,4 Lic. Carlos M. Fernández Andreu,5 Lic. Mayda R. Perurena Lancha6 y Téc. Dianeya Mendoza Llanes7
Se propuso la obtención de antisueros en el Laboratorio de Micología del Instituto de Medicina Tropical Pedro Kourí, que permitiera la rápida identificación de las variedades y serotipos de Cryptococcus neoformans, dada la importancia clínico-epidemiológica que reviste su conocimiento. Mediante la inmunización de conejos con células enteras formalinizadas de los 4 serotipos de esta levadura, se obtuvieron antisueros que fueron adsorbidos con células de los serotipos heterólogos; lográndose 3 antisueros capaces de diferenciar a la var. neoformans (anti AD) y a sus respectivos serotipos (anti-A y anti-D). Este trabajo contribuirá al conocimiento de la epidemiología de esta importante enfermedad en el medio cubano, por ser la primera vez en Cuba que se logra la obtención de antisueros útiles para la identificación de los serotipos pertenecientes a la variedad neoformans, principal agente causal de la criptococosis.
Palabras clave: Cryptococcus neoformans, criptococosis, epidemiología, sueros inmunes.
Atendiendo a la cantidad de manosa y la extensión de la O-acetilación así como otras propiedades bioquímicas de la cápsula polisacarídica, las cepas de Cryptococcus neoformans pueden ser agrupadas en 2 variedades y estas a su vez en serotipos: C. neoformans var. neoformans (serotipos A y D) y C. neoformans var. gattii (serotipos B y C).1,2 Se han descrito diferentes métodos de serotipificación entre los cuales se destaca Crypto Check Iatron desarrollado por Ikeda y otros en 1982 basado en la técnica de aglutinación en lámina.3
Estudios realizados previamente en el laboratorio del Instituto de Medicina Tropical "Pedro Kourí" utilizando medios de cultivo como el L-canavanina-glicina-azul de bromotimol, han demostrado la prevalencia de la variedad neoformans en Cuba;4 no obstante, solo se ha podido llevar a cabo la serotipificación de cepas procedentes de aislamientos clínicos.5
Dado el incremento de la criptococosis en las últimas décadas y la importancia que reviste la identificación de los serotipos en el conocimiento de la posible fuente de infección, los autores se propusieron el desarrollo de un método rápido que permitiera su identificación en el laboratorio.
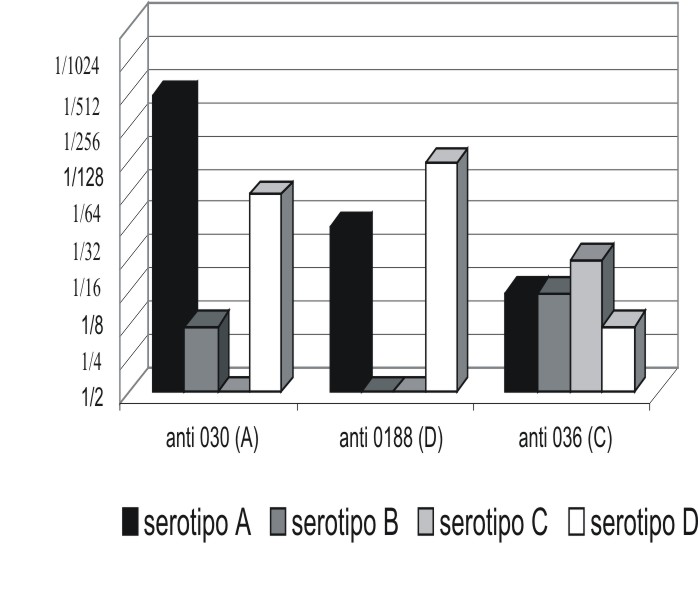
]]> Para la obtención de antisueros específicos se siguió la metodología descrita por Illnait y otros.6 Las suspensiones antigénicas fueron preparadas con células enteras formalinizadas a partir de cepas ampliamente estudiadas y caracterizadas en el laboratorio (LMIPK 030 serotipo A, LMIPK 0188 serotipo D, LMIPK 0186 serotipo B y LMIPK 036 serotipo C. Se emplearon conejos de la raza Nueva Zelandia de 3 kg de peso procedentes del Centro Nacional para la Producción de Animales de Laboratorio, Cuba. Los sueros obtenidos fueron titulados mediante aglutinación en lámina utilizando cepas homólogas. Para determinar el grado de reacción cruzada con otros serotipos se procedió de igual manera pero con serotipos heterólogos (fig.).Fig. Título de aglutinación de los antisueros obtenidos con las cepas homólogas y heterólogas.
Todos los antisueros obtenidos mostraron reacción cruzada con las cepas heterólogas. Estas fueron más evidentes entre los serotipos A y D, posiblemente originadas por la similitud estructural de ambas cápsulas.7 Por su parte el antisuero antiserotipo C manifestó fuerte reacción con los restantes serotipos, lo que pudiera relacionarse con la mayor complejidad en la estructura de su envoltura (fig.).7
Trabajos realizados por Gade y otros7 y Murphy8 demostraron que los serotipos A y D son inmunológicamente más activos, resultados que se repiten en este trabajo. En el caso de los conejos inmunizados con la cepa LMIPK 0186 (serotipo B) no se logró respuesta de anticuerpos evidente; lo que pudiera haber estado sujeto a la capacidad de respuesta individual de los animales inmunizados o a la pobre inmunogenicidad de la cepa empleada.
Las reacciones cruzadas fueron eliminadas mediante adsorción5 y dilución seriada de los sueros.9 Después de aplicados estos procedimientos, los niveles de anticuerpos resultantes en el antisuero anti-036 (serotipo C), no fueron suficientes para su uso en la caracterización serológica (datos no mostrados).
Finalmente fueron obtenidos 3 antisueros capaces de reconocer a la variedad neoformans: anti AD, anti-A y anti-D, los cuales fueron enfrentados a cepas controles previamente caracterizadas, demostrándose su confiabilidad (tabla).
Tabla. Caracterización serológica de las cepas de C. neoformans y especies relacionadas
| Cepas | ]]> Antisueros | Resultados | ||
| Anti AD | Anti D | Anti A | ||
| C. neoformans LMIPK 030 (A) | (+) | (-) | (+) | A |
| C. neoformans 235 (A) | ]]> (+) | (-) | (+) | A |
| C. neoformans LMIPK 0100 (A) | (+) | (-) | (+) | A |
| C. neoformans LMIPK 0193 (A) | ]]> (+) | (-) | (+) | A |
| C. neoformans LMIPK 0135 (B) | (-) | (-) | (-) | no aglutinó |
| C. neoformans LMIPK 0186 (B) | ]]> (-) | (-) | (-) | no aglutinó |
| C. neoformans LMIPK 024 (B) | (-) | (-) | (-) | no aglutinó |
| C. neoformans LMIPK 036 (C) | ]]> (-) | (-) | (-) | no aglutinó |
| C. neoformans LMIPK 025 (C) | (-) | (-) | (-) | no aglutinó |
| C. neoformans LMIPK 026 (D) | ]]> (+) | (+) | (-) | D |
| C. neoformans LMIPK 0188 (D) | (+) | (+) | (-) | |
| Candida albicans | (-) | ]]> (-) | (-) | no conocido |
Por causa de que los estudios hasta el momento realizados tanto con muestras clínicas como ambientales en la región, y muy especialmente en Cuba, han demostrado el predominio del serotipo A, y a la difícil disponibilidad de los reactivos comerciales para la serotipificación de las cepas de C. neoformans, es de considerar como un gran paso de avance la obtención de antisueros que permitan un mejor conocimiento de la criptococosis en Cuba.
The obtention of antisera at the Mycology Laboratory of Pedro Kouri Institute of Tropical Medicine was proposed to allow the rapid identification of the varieties and serotypes of Cryptococcus neoformans, since it is very important from the clinicoepidemiological point of view to know them. By immunizing rabbits with formalized whole cells of the 4 serotypes of this yeast, it was possible to obtain antisera that were absorbed with cells of the heterologous serotypes. 3 antisera capable of differentiating the neoformans var. (anti AD) and its respective serotypes (anti-A and anti-D) were attained. This paper will contribute to the knowledge of the epidemiology of this important disease in the Cuban setting for being the first time that useful antisera are obtained in Cuba to identify the serotypes corrresponding to the neoformans variety, which is the main causal agent of criptococcosis.
Key wods: Cryptococcus neoformans, cryptococcosis, epidemiology, immune sera.
Recibido: 23 de junio de 2003. Aprobado: 2 de agosto de 2003.
Dra. María T. Illnait Zaragozí. Instituto de Medicina Tropical Pedro Kourí. Apartado 601, Marianao 13, Ciudad de La Habana, Cuba. Correo electrónico: mtilnait@ipk.sld.cu
1 Máster en Bacteriología-Micología. Especialista de II Grado en Microbiología. Investigadora Agregada.
2 Licenciado en Microbiología. ]]>
3 Máster en bacteriología-Micología. Licenciada en Microbiología. Aspirante a Investigadora.
4 Especialista de II Grado en Microbiología. Investigador Agregado.
5 Máster en Microbiología. Licenciado en Microbiología. Investigador auxiliar.
6 Máster en Bacteriología-Micología. Licenciada en Microbiología. Investigadora Agregada.
7 Técnica en Microbiología.