TECHNIQUE
Evaluation of four viroid RNA extraction methods for the molecular diagnosis of CEVd in Citrus lemon using RT-PCR, Dot blot and Northern blot
Evaluación de cuatro métodos de extracción de ARN viroide para el diagnóstico molecular de CEVd en Citrus limon mediante RT-PCR, Dot blot y Northern blot
]]>
Rodolfo Umaña1, Clara Pritsch1, Juan R Arbiza2, Fernando Rivas3, Gabriela Pagliano1
1 Departamento de Biología Vegetal, Facultad de Agronomía, Universidad de la República. Garzón 780, Montevideo, Uruguay.
2 Sección Virología, Facultad de Ciencias, Universidad de la República. Iguá 4225, Montevideo, Uruguay.
3 Instituto Nacional de Investigación Agropecuaria, INIA. Ruta 3 Camino al Terrible. Salto, Uruguay.
ABSTRACT
An efficient method for RNA extraction that leads to RNA high yield and purity is a technical issue relevant for development and optimization of molecular diagnostic methods aimed to detect viroid infections in citrus varieties. Residual contaminants may affect RNA detection depending on the molecular diagnosis approaches. This condition can be evaluated through RNA absorption spectrum analysis. Functionally, it is assessed through observation of RT-PCR amplification products and Northern blot and Dot-blot signal intensities, displaying levels of analytical response/sensitivity. Four RNA extraction methods were evaluated to determine their effects on the capacity to detect viroid CEVd presence/absence in Citrus limon through four molecular diagnostic approaches: 1) conventional viroid extraction (CVE); 2) phenol/guanidine thiocyanate (PGT), 3) SDS/potassium acetate (SPA); and 4) formaldehyde/ SSC (FS). Phloem tissue quantifications showed values between 7500 ng/µL and 1200 ng/µL and ranged 1.3-2.0 OD260/280. Evaluations through RT-PCR showed the expected amplifications of the entire CEVd genome, but erratic scenarios still remained. Non-radioactive probe hybridization techniques revealed high intensity signals (132 RU) for infected tissue, by using the CVE method, and a positivity cut-off for the presence of infection was established (78 RU). Nevertheless, molecular hybridization tools can jeopardize the diagnosis due to the thoroughness of the protocol and the RNA template conditions. The diagnostic ability of the association of Northern blot with CVE viroid extraction analyses as starting point was evidenced for successful detection, among the molecular methods tested.
]]> Keywords: CEVd, diagnostic, non-radiactive Northern blot, RT-PCR, citrus viroids.
RESUMEN
En el desarrollo de metodologías moleculares diagnósticas en variedades cítricas propensas a infección viroide, se precisa la extracción eficiente de ARN, siguiendo criterios de concentración y pureza. Los contaminantes remanentes pueden afectar la detección según la herramienta molecular escogida. Esta condición se analiza a partir del espectro de absorción del ARN; mediante la reacción en cadena de la polimerasa con transcriptasa inversa (RT-PCR) y por los niveles de intensidad de las señales de Northern blot y Dot blot, en términos de respuesta analítica/sensibilidad. Se evaluaron cuatro métodos de extracción de ARN, por sus efectos sobre la detección de presencia/ausencia del Citrus Exocortis Viroid (CEVd) en Citrus limon mediante análisis moleculares diagnósticos: 1) extracción viroide convencional (EVC); 2) fenol/tiocianato de guanidina (FTG); 3) SDS/acetato de potasio (SAP); y 4) formaldehído/SSC (FS). Los valores del tejido de floema estuvieron entre 7500 ng/µL y 1200 ng/µL y los rangos entre 1.3 y 2.0 DO
Palabras clave: CEVd, diagnóstico, Northern blot no radiactivo, RT-PCR, viroides cítricos.
INTRODUCTION
Citrus viroids (CVd) are subviral pathogenic entities of the Pospiviroidae family. They lack the capsid protein and are formed solely by circular RNA (284-375 nucleotides), a covalently closed non-coding single strand with regions of high intramolecular complementarity [1].
]]> According to the classification suggested by Flores et al. [2] with the modifications of the Viroid Study Group of the International Committee on Taxonomy of Viruses (ICTV) (www.ictvonline.org/virusTaxonomy.asp), the CVd are formed by the following species: CEVd, CBLVd, HSVd, CDVd, CBCVd, CVd-V and CVd-VI.In general, the diagnosis of CVd infections involves important limitations. The methods based on biological assays are cumbersome, hardly specific, comprise the amplification of the viroid in indicator plants and are time-consuming. On the other hand, molecular diagnosis methods based on the detection of complete viroid genomes are affected by CVd secondary structure as well as by CVd interactions with various molecules of the infected tissue.
The absence of capsid has prevented the use of immunological techniques, while the strong intramolecular pairing and the low and irregular titers of viroid particles in infected tissues have led to mistaken molecular diagnoses [3, 4].
Several molecular detection methods of CVd have been developed in order to increase sensitivity, specificity and precision of results, decrease diagnostic time and optimize the use of low toxic, non-radioactive products. These methods are based on either the enzymatic amplification of the viroid genome (by reverse transcriptase-polymerase chain reaction; RT-PCR) or on RNA-RNA or RNA-DNA hybridizations. They also share the requirement for simple and inexpensive RNA extraction protocols to obtain plant nucleic acids preparations enriched with viroid RNA (vdRNA), and with levels of contaminants as low as possible within the RNA preparation to avoid interferences on the detection process [5-7].
The RNA extraction protocols for the detection of viroids [8-11] differ on the type of plant (field-grown or bioindicator) or tissue sampled, chemical treatment of the sample, length of handling period, level of vdRNA recovery and the amount of residual contaminants. As a consequence, the RNA extraction method must be chosen and adjusted according to which detection method will be followed. The polymerase chain reaction (PCR) is widely used for the diagnosis of citrus viroids [12-14]. Although the reverse transcriptase-PCR (RT-PCR) of the viroid genome has been successfully used for cloning and sequencing viroids [15], specificity and efficiency limitations have been described. These include the great similarity of sequences with the viroid genomes of other genera, making the design of specific probes difficult [16, 17]. Moreover, the secondary structure determines the presence of amplicons of unexpected sizes having the potential for renaturing and restructuring the viroid sequence at the lower temperatures of the RT-PCR [18]. The permanence of inhibitors derived from RNA extraction, such as phenol compounds and polysaccharides, may produce false negatives. The unspecific amplification of high and low molecular weight PCR products from field samples is frequent, with confusing results affecting the reliability of the diagnosis [15, 19]. As a consequence, the inclusion of RT-PCR as a routine tool in certification and quarantine programs has been questioned [6, 20].
Molecular hybridization is another strategy for the detection of viroids, which depends on a successful association of the probe with the vdRNA used. Hence, the treatment of vdRNA with heat and denaturing reagents is important in performing the technique [7]. The probes of complementary DNA (cDNA) marked with digoxygenin (DIG-11-UTP) are frequently used because they are easy to handle, of rapid synthesis, non-radioactive, do not require special equipment or protection, and are non-contaminants. The Dot blot hybridization technique is a rapid diagnostic tool that allows to handle massive samples, being an attractive diagnostic choice for sanitation and certification programs in citrus [5, 9]. However, the use of Dot Blot for direct viroid detection in field-grown plants (without a first amplification passage in Etrog citron) has produced errors and false positive results. For example, under low astringency conditions and in the abscense of denaturing agents, false positives as well as confusing results associated to visual detection are frequently present [21, 22]. The observations of WenXing et al. [23] revealed the possible interactions of viroid DNA probes with host ribonucleoprotein complexes, producing erratic signals in Dot blot molecular hybridizations. Gómez and Pa-llás [24] demonstrated the formation of in vivo RNA-protein complexes between the HSVd and the lectin protein 2 in cucumber phloem (the most abundant protein in phloem). These proteins may produce the interference or binding of the probes in uninfected samples that are analyzed using RNA Dot blot. The viroids may suffer modifications in the molecular geometry of their structural motifs and potentiate interactions with plant proteins [25]. This complex may obstruct the pairing of the marked probe with the genome of the pathogen. Hence, the Dot Blot diagnostic technique is not recommended to be applied as the sole diagnostic method of viroids when studying field-grown citrus plants [11, 23].
According to Murcia et al. [7], diagnostics by Dot blot hybridization can detect small amounts of vdRNA in the infected plant material. Using the appropriate cDNA probes (DIGcDNA), the technique enables the discrimination of known CVd species and the generation of reliable and consistent results. These authors proposed this molecular hybridization strategy for sanitation programs, phytosanitation improvement, quarantine and certification, because of its high sensitivity and efficiency. In this work, four vdRNA extraction protocols were compared on their efficiency to recover citrus viroid particles and their effect on viroid particles detection by three molecular diagnostic tools.
MATERIALS AND METHODS
RNA extraction methods
]]> Tissue samples from Citrus limon (L.) Burm. (Lisbon variety) with accession numbers K395 and CDL384 were used as positive and negative controls of the infection with Citrus Exocortis Viroid (CEVd), respectively, according to previous studies [26]. Control plants were grown in a commercial orchard in Uruguay. Two other positive and negative controls were included, comprising tissue sampled from Etrog citron infected with CEVd isolate 17 (CEVd-17) and Etrog citron without infection, respectively. Citron plants were grown under controlled conditions in a growth chamber of the Biotechnology Laboratory of the Agronomy Faculty, Universidad de la República.For RNA extraction, the bark of branches 0.1 to 1 cm in diameter (where the phloem tissue is locat-ed) were pulverized with liquid nitrogen using a modified domestic grinder (UfesaTM). The amount of pulverized tissue in all extraction methods was standardized at 5 g and nucleic acids were resuspended in 300 μL of double-deionized sterile water. The extraction methods of vdRNA assessed for lemon tissue samples were:
CVE
Conventional viroid extraction method (CVE), designed to obtain a high viroid titers from the specific plant material [27]. The pulverized tissue was homogenized with extraction buffer (0.4 M Tris-HCl, pH 8.9; 1 % (w/v) SDS; 5 mM EDTA, pH 7.0; 2 % (v/v) mercaptoethanol) containing phenol saturated in water at a neutral pH and facilitated by shaking in a vortex. Following centrifugation at 8000 × g, for 20 min (at 4 °C), the aqueous phase was recovered and treated with one tenth of the volume of 3 M sodium acetate pH 5.5, and 3 volumes of cold absolute ethanol. It was incubated for 1 h at -20 °C (the precipitation of total nucleic acids was evident by the flocculation of the solution) and centrifuged at 8000 × g, for 20 min (at 4 °C). Total nucleic acids were dialyzed using dialysis tubes (Sigma-Aldrich; 33 mm, cellulose membrane with retention of molecules of more than 12 400 MW) in a TKM 1× solution (10 mM Tris-HCl, pH 7.4; 10 mM KCl; 0.1 mM MgCl2). The dialyzed preparation was partitioned in 2 M LiCl and the soluble fraction was concentrated by precipitation with absolute ethanol.
PGT
The phenol/guanidine thiocyanate method (PGT) is based on the steps described by Chomczynski and Sacchi [28]. First, the pulverized tissue was homogenized in 15 mL of the TriPure® reagent (Roche™). The homogenate was then clarified through centrifugation, the supernatant was collected and phases were separated with 0.2 mL of chloroform per milliliter of TriPure®. Finally, RNA was isolated from the aqueous phase through precipitation with 0.5 mL of isopropanol per milliliter of TriPure® and washed with 75 % ethanol.
SPA
The Sodium Dodecyl Sulfate (SDS)/potassium acetate method (SPA), reported by Cañizares et al. [29], was used with certain modifications. First, five grams of tissue were homogenized in the extraction buffer (0.1 M Tris-HCl, pH 8.9; 50 mM EDTA; 0.5 M NaCl; 25 mM mercaptoethanol). Next, the resulting solution was treated with 20 % SDS (pH 7.2) and incubated at 65 °C for 20 min, and 5 M potassium acetate (pH 7.5) was further added, followed by incubation for 30 min on ice. After that, the phases were separated by centrifugation (15 min at 12 000 × g, 4 °C); the supernatant was collected and equal volumes of a 20 % polyethylenglycol solution and 1 M NaCl were added. This was left to stand for 1 h at 4 °C, and was subsequently centrifuged at 12 000 × g for 15 min at 4 °C, in order to isolate the high molecular weight RNA and recover small RNAs in the supernatant. Finally, a precipitation step was carried out with absolute ethanol.
FS
The formaldehyde/saline sodium citrate (SSC) buffer method (FS) was originally implemented for the molecular diagnosis of the PSTVd viroid [30]. Two mL of the extraction buffer (5× SSC / 18.5 % formaldehyde) were added per gram of pulverized tissue. The solution was homogenized using a vortex shaker with 0.5 volumes of saturated phenol in water (pH 7.0) and 0.5 volumes of chloroform, followed by centrifugation at 5000 × g for 5 min, at 4 °C. The supernatant was then collected and nucleic acids were precipitated with isopropanol through incubation at -20 °C for 1 h. It was later centrifuged at 3000 × g, for 30 min at 4 °C, and washed with 75 % ethanol. RNA was extracted from infected and uninfected citron control tissues by using the CVE method.
]]> Nucleic acid quantificationConcentrations of plasmid DNA (pDNA), probes-DNA:DIG, total RNA of citron leaves and RNA from the phloem tissue of C. limon were measured by UV-visible spectrophotometry (NanoDrop 1000, Thermo Scientific), and data on concentration were obtained in nanograms per microliter. Samples purity was estimated by the OD260/280 coefficient, regarded as optimal for values higher than 1.9 for RNA and 1.8 for DNA [31].
RT-PCR
The reactions were carried out in a PALM PCR cycler (Corbett Research). The synthesis of the first strand of the CEVd viroid was done with the complementary primer (CEVd-R1) 5’-CCGGGGATCCCTGAAGGA- 3’ [32]. The RNA template (~ 200 ng) obtained from each extraction method was denatured in a first step at 85 °C for 5 min; later at 55 °C, for 1 min in 1.25 μM of the CEVd-R1 primer (final volume of 8 μL), followed by an incubation on ice for 3 min. In the second step of the reaction, the first strand of the viroid cDNA was synthesized by using 50 U of RevertAid™ reverse transcriptase M-MuLV RNase H-(Fermentas) in a reaction mixture (50 mM Tris-HCl, pH 8.3; 4 mM MgCl2; 50 mM KCl; 10 mM DTT (Fermentas); 0.4 mM of each dNTP) with a final volume of 13 μL, and incubated at 42 °C for 1 h, followed by a 3-min enzymatic inactivation at 90 °C. The second strand of viroid cDNA was synthesized in a mixture with a final volume of 25 μL (4 μL of the reaction of the first strand, 0.5 U of TaqDNA polymerase (Invitrogen®); 20 mM Tris-HCl, pH 8.4; 50 mM KCl, 1.6 mM MgCl2; 0.2 mM dNTP and 0.5 μM of primers CEVd-R1 5’-CCGGGGATCCCTGAAGGA-3’ and CEVd-F1 5’-GGAAACCTGGAGGAAGTCG-3’ [32]. Amplification conditions were: denaturing at 94 °C for 5 min, followed by 36 cycles (at 94 °C for 30 s, at 55 °C for 30 s and 72 °C for 1 min) and a final cycle at 72 °C for 5 min.
Preparation of PCR-labeled probes
The cDNA corresponding to the complete genome of the CEVd viroid was labeled through PCR by incorporating digoxigenin (DIG-11-dUTP; Roche Molecular Biochemicals) [33]. The reaction mixture included 225 pg of the recombinant plasmid pGEM®-T-Easy-CEVd clone 054.11 in a solution composed of 20 mM Tris-HCl (pH 8.4), 50 mM KCl, 1.6 mM MgCl2, a mixture of 0.2 mM dNTPs (GTP, CTP, ATP), 0.14 mM TTP, 0.5 U of TaqDNA polymerase (Invitrogen®), 1 nmol of alkaline-stable DIG-11-dUTP, and each one of the complementary primers CEVd-R1 and the hom-ologue CEVd-F1 at 0.5 μM [34]. The amplification conditions are the same as described in the synthesis of the second strand of the cDNA.
Non-isotopic molecular hybridization
For the Northern blot analysis, the RNA extracted by each of the methods (aliquots of 20 μL corresponding to 333 mg of fresh tissue weight) was separated by electrophoresis (a vertical structure with glass sheets 10 × 17 cm) in 5 % polyacrylamide gels, buff-er TAE 1×, under non-denaturing conditions, for 3 h at 60 mA. The complex of total nucleic acids was visualized in an ultraviolet transilluminator (UV) after ethidium bromide staining. In order to approximately identify the gel region containing the vdRNA, a fragment of the gel was recovered that included the 5S ribosomal RNA (5S rRNA) and the signal recognition particle 7S RNA, considering an upper margin of 1 cm and a lower margin of 2 cm, vertically. The gel segment was subjected to electrotransference (400 mA, buffer TBE 1× for 1.5 h) to positively-charged Nylon membranes (Roche Applied Science) [33].
For the Dot blot assays, the RNA of each one of the extraction methods (6 μL equivalent to 100 mg of fresh tissue weight) were pre-treated with 6 μL of formamide, 1.5 μL of 20× SSC and incubated for 15 min at 68 °C [34]. The sample was dotted in Nylon+ membranes (Roche Applied Science) and dried at room temperature (RT). In all hybridization trials, membranes were exposed to a treatment consisting of 70 000 μJ/cm2 for UV crosslinking in a hybridization oven (Hoefer-Uvc500, Amersham Biosciences Corp.) to fix the nucleic acids. The pre-hybridization (42 °C, 2 h) and hybridization (60 °C over night) steps were carried out in a solution containing 50 % formamide, 5× SSC buffer (150 mM NaCl, 15 mM sodium citrate; pH 7.0) with 0.02 % SDS, 0.1 % N-laurylsarcosin and 2 % of a blocking solution (w/w; Roche Applied Science) [7], to destabilize the hydrogen bonds of the immobilized nucleic acid strands, and therefore, to avoid the typical intra-molecular pairing of CVd. Before hybridization, the marked probes were denatured (690 ng) for 5 min at 95 °C and immediately placed on ice. After hybridization, the membranes were washed twice in SSC 2×, 0.1 % SDS at RT for 15 min and then incubated at 60 °C for 1 h in SSC 0.1×, 0.1 % SDS. Finally, they were washed with washing buffer (0.1 M maleic acid, 0.15 M NaCl, pH 7.5 and 0.3 % (v/v) Tween 20), for 5 min at RT. The membrane was blocked with 1× blocking reagent (Roche Applied Science) for 40 min at RT. The probe-DIG:vdRNA hybrids were detected with a anti-DIG Fab fragment-alkaline phosphatase conjugate, at 0.3 U of antibody per milliliter of the 1× blocking solution. The nonspecifically-bound antibody was washed out of the membrane twice with a washing buffer solution, for 15 min at RT, followed by stabilization with the detection buffer (0.1 M Tris-HCl, 0.1 M NaCl; pH 9.5) for 30 min. Finally, the vdRNA-DIG-Fab-alcaline phosphatase hybrids were visualized by including the chemiluminescent substrate CSPD at 0.35 mM (Roche Applied Science) through incubation for 30 min, and X-ray films were exposed for 20 min, at 37 °C. The visible light emitted by CSPD dephosphorylation was retained by the films, which were developed by autoradiography. The hybridization signals produced were quantified by densitometry analysis of the developed plate using the 1 DScanEX program version 3.1 Demo (Scanalytics, Inc.). An image-estimated background correction method was used (automatic correction with minimum pixels). The intensities of the signals were reported in relative units (RU). The graphic representation of columns was done with the InfoStat© program version 2008 student.
]]> RESULTS AND DISCUSSION
At first, four RNA extraction protocols were compared in terms of values for RNA concentration and purity, based on bark tissues sampled from C. limon plants collected in the field. The CVE and PGT methods produced RNA samples with the highest purity levels (DO260/280 ranging from 1.9 to 2.0), with low levels of contaminants sensitive to UV as proteins and phenol. These two methods also showed the highest mean total RNA concentrations: 2330 ng/μL and 7650 ng/μL, respectively. In contrast, the SPA and FS extractions recovered low purity RNA (DO260/280 ranging from 1.3 to 1.5), with RNA concentrations of 1217 ng/μL and 2400 ng/μL, respectively. In general, good RNA yield recovery has been reported when phloem tissue is sampled, since viroids are mostly transported by the phloem [35]. Similar results reported for Etrog citron indicate that the concentration of the CEVd viroid in the bark is 10 times higher than in the leaves [36]. Nevertheless, the high viroid titer detected in the foliar tissue of Etrog citron is explained by the high bio-amplification of the pathogenic genome in this sensitive host.
The high values of RNA concentration observed with CVE, PGT and FS methods may be due to the fact that these methods included a phenol treatment that led to an efficient disruption of cell membranes as well as removal of proteins in the organic phase. Nevertheless, it is likely that the high RNA yield with PGT may be related to the absence of steps to recover mostly low molecular weight RNA rather than total host RNA derives in an overestimation of RNA in the sample analyzed. It must be pointed out that the purity of RNA obtained by FS was much lower than that obtained by CVE. This may be caused by the absence in the FS method of differential purification steps such as: dialysis and LiCl precipitation. Finally, the low concentration and purity of the RNA recovered through the SPA method is possibly due to losses in the recovery of the RNA in the differential polyethylene glycol precipitation steps, the absence of steps to capture or discriminate contaminants released in the pulverization of the material with liquid nitrogen, or both.
Subsequently, the possible detrimental effect of the RNA extraction method on the amount of detectable CEVd was analyzed by three diagnostic techniques: RT-PCR, Northern blot and Dot blot. For this, we analyzed positive (K395) and negative samples (CDL384) for CEVd, of C. limon (v. Lisbon) plants collected in the field, according to the preceding results [36].
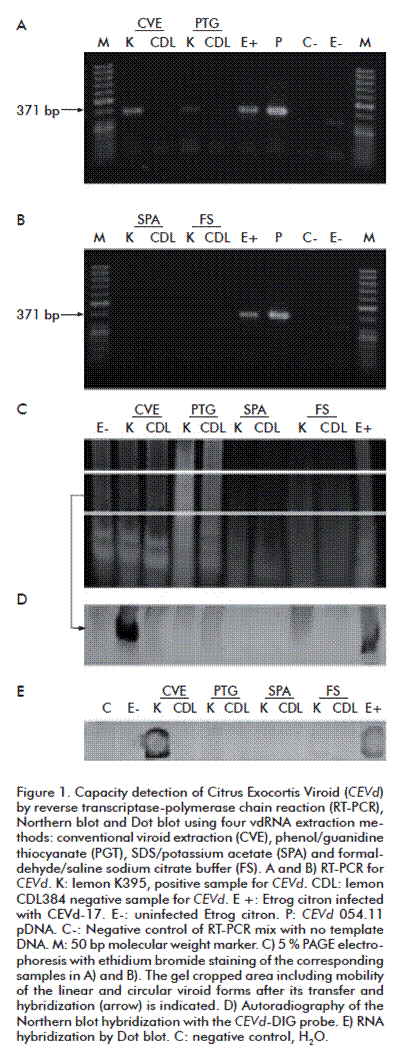
In RNAs obtained by the CVE and PGT methods, evidences were observed of total genome amplification of CEVd (371 pb) by RT-PCR from the K395 lemon plant (Figure 1A). This is probably due to the high level of purity in the sample, which relates to reduced concentrations of RT-PCR inhibitors such as polysaccharides and polyphenols, which are typical of the cell wall of citrus tissues. However, because the RNA analyzed by RT-PCR was highly diluted (approximate amounts of 200 ng) it could also be possible that the effect of sample’s dilution would have decreased the concentration of potential contaminants which interfere in viroid detection. The extractions with the SPA and FS methods did not show amplicons in the RT-PCR assay (Figure 1B), which would be due to an excess of enzymatic activity inhibitors in the sample. These findings are in contrast to observations by Bernard and Duran-Vila [6], who indicated that the quality of the RNA obtained from the SPA extractions without modifications made the analysis by RT-PCR feasible. Perhaps, the differences in the extraction protocol, reaction conditions, primers, and enzymes used may explain this discrepancy [6, 37].
In the negative control, the uninfected Etrog citron showed amplicons with lower than expected molecular weights (371 bp; Figures 1A and B, lane E-). Mixed populations of high and low molecular weight amplicons would be produced by unspecific hybridization, which occurs during the interaction of the reverse transcriptase with host RNA templates bearing thermostable hairpins or by the effect of native RNAs that fold back on themselves generating a primer for the reverse transcriptase [38]. Finally, the formation of secondary structures in the viroid genome at low reaction temperatures may limit the access of the enzyme and produce smaller amplicons than those expected [15].
The molecular hybridization of viroid RNA by Dot blot clearly detected the CEVd viroid in the K395 positive sample obtained through the CVE method, while no clear signals were detected when using other extraction methods (Figure 1E). The negative control samples showed no signals. These results would indicate that the ability to detect the RNA sequence of the viroid genome by hybridization with the double-stranded DNA-DIG probe is affected by the extraction method used.
The detection of the CEVd in C. limon using Northern blot-CEVd-DIG showed a clear (high level of saturation, 132 RU) and compact signal on a radiographic plate (Figure 1D) solely for the RNA extracted by the CVE method (lane K of CVE). In contrast, we did not observe clear evidence of the presence of CEVd when this infected tissue was processed by the PGT, SPA or FS (~ 30 RU) extraction methods. The high signal intensity observed only for CVE extractions up to a saturation point may be explained because there was an increase in the titer of the viroid fixed on the membrane and paired with the CEVd-DIG probe (Figures 1D and E, and Figure 2).
]]> The low intensity of molecular hybridizations with the other three RNA extraction methods may originate from reduced efficiency of probe labeling. In this case, the pairing kinetics between the probe and the vdRNA would be limited by the high number of copies of the homologous sequence of RNA-CEVd paired with unlabeled complementary probes. However, in this study the labeling efficiency was high, since saturation was observed in the intensity of the Northern blot signal with a minimum amount of the labeled probe (690 ng). These results are in agreement with those of Murcia et al. [7], since the CVE followed by Northern blot detection was described as reliable and robust to detect viroids in field material.The lower sensitivity of the molecular hybridization compared to RT-PCR is highlighted, since the PGT extraction does not produce compact signals in the ra-diographic plate (Figure 1D and E), but reveals the presence of the viroid in the RT-PCR (Figure 1A and B).
Similar to results using RNA Dot blot, the extractions using SPA, PGT and FS showed significantly lower signal intensities than those obtained with CVE in Northern blot hybridizations, in infected lemon RNA and the citron CEVd-17 positive control (Figure 2). The quantification of these signals with the specific probe CEVd-DIG in phloem tissue from C. limon shows a possitivity cut-off signal (78 RU) and infection (according to the parameters for background correction estimated by the image). This is based on the strict criterion that an infected sample must have a signal twice the highest intensity value of the negative control sample (39 RU), in order to establish an acceptance interval for a positive signal and to provide certainty on the presence of a detectable viroid titer in the sample [39].
Overall, the results of this study showed that the differences in concentration and purity of the RNA obtained through the four extraction methods affected the levels of vdRNA detectable through three molecular diagnostic techniques. Using molecular detection tools based on enzymatic amplification reactions (RT-PCR) and hybridization with labeled cDNA probes, we demonstrated that under our experimental conditions using infected tissues of citrus varieties in the field, the CVE method recovered the largest amount of copies of vdRNA with high levels of purity.
Previous reports had recommended the use of Northern blot analysis in the detection of CVd in citrus varieties growing in commercial orchards, which avoids the need of recurring to a previous bio-amplification passage of the viroid in indicator plants [7, 33]. Based on our results, we claim the CVE extraction method followed by Northern blot approach as the most appropriate test for systematic screening of citrus plants including propagation materials within sanitation programs, and in eventual prospective sanitations for the control and eradiation of the citrus viroids described.
ACKNOWLEDGEMENTS
We would like to thank BID-CONICYT for the funding project PDT 74/19; Dr. Nuria Duran-Vila for her support and recommendations in this study; Agr. Eng. Jacques Borde and Agr. Eng. Ana Bertalmío for providing the plant tissues used as controls for this study; MSc. Paola Gaiero for critically reviewing the manuscript.
]]> REFERENCES
1. Flores R. A naked plant-specific RNA ten-fold smaller than the smallest known viral RNA: the viroid. C R Acad Sci III. 2001;324(10):943-52.
2. Flores R, Randles JW, Bar-Joseph M, Diener TO. Viroids. In: van Regenmortel MHV, Fauquet CM, Bishop DHL, Carsten EB, Estes MK, Lemon SM, et al., editors. Virus taxonomy. Seventh Report of the International Committee on Taxonomy of Viruses. San Diego: Academic Press; 2000. p. 1009-24.
3. Palacio A, Foissac X, Duran-Vila N. Indexing of citrus viroids by imprint hybridisation. Eur J Plant Pathol. 1999;105(9):897-903.
4. Barbosa CJ, Pina JA, Navarro L, Duran-Vila N. Replication/accumulation and symptom expression of citrus viroids on some species of citrus and related genera. In: Duran-Vila N, Milne RG, Da Graça JV, editors. Proceedings XV International Conference of the Organization of Citrus Virologists (IOCV). Riverside, CA: International Organization of Citrus Virologists; 2002. p. 264-71.
5. Cohen O, Batuman O, Stanbekova G, Sano T, Mawassi W, Bar-Joseph M. Construction of a multiprobe for the simultaneous detection of viroids infecting citrus trees. Virus Genes. 2006;33(3):287-92.
6. Bernard L, Duran-Vila N. A novel RT-PCR approach for detection and characterization of citrus viroids. Mol Cell Probes. 2006;20(2):105-13.
7. Murcia N, Serra P, Olmos A, Duran-Vila N. A novel hybridization approach for detection of citrus viroids. Mol Cell Probes. 2009;23(2):95-102.
8. Nakahara K, Hataya T, Uyeda I. A simple rapid method of nucleic acid extraction without tissue homogenization for detecting viroids by hybridisation and RT-PCR. J Virol Methods. 1999;77(1):47-58.
9. Noronha-Fonseca ME, Marcellino LH, Gander E. A rapid and sensitive dot-blot hybridization assay for the detection of citrus exocortis viroid in Citrus medica with digoxigenin-labelled RNA probes. J Virol Methods. 1996;57(2):203-7.
10. Palacio A, Foissac X, Duran-Vila N. Indexing of citrus viroids by imprint hybridization: comparation with other detection methods. In: Da Graça JV, Lee RF, Yokomi RK, editors. Proceedings XIV Conference of the Internacional Organization of Citrus Virologist (IOCV). Riverside, CA: Internacional Organization of Citrus Virologist; 2000. p. 294-301.
11. Ito T, Ieki H, Ozaki K. Simultaneous detection of six citrus viroids and Apple stem grooving virus from citrus plants by multiplex reverse transcription polymerase chain reaction. J Virol Methods. 2002;106(2):235-9.
12. Ragozzino E, Faggioli F, Barba M. Development of a one tube-one step RT PCR protocol for the detection of seven viroids in four genera: apscaviroid, hostuviroid, pelamoviroid. J Virol Methods. 2004;121(1):25-9.
13. Wang X, Zhou C, Tang K, Zhou Y, Li Z. A rapid one-step multiplex RT-PCR assay for the simultaneous detection of five citrus viroids in China. Eur J Plant Pathol. 2009;124(1):175-80.
14. Tessitori M, Rizza S, Reina A, La Rosa R. Development of a real-time assay for the simultaneous detection of citrus viroids [abstract]. J Plant Pathol. 2004;86(4 Special issue):336.
15. Navarro B, Darós JA, Flores R. Reverse transcription polymerase chain reaction protocols for cloning small circular RNAs. J Virol Methods. 1998;73(1):1-9.
]]>16. Puchta H, Ramm K, Luckinger R, Hadas R, Bar-Joseph M, Sänger HL. Primary and secondary structure of citrus viroid IV (CVd IV), a new chimeric viroid present in dwarfed grapefruit in Israel. Nucleic Acids Res. 1991;19(23):6640.
17. Rakowski AG, Szychowski JA, Avena ZS, Semancik JS. Nucleotide sequence and structural features of the group III citrus viroids. J Gen Virol. 1994;75(Pt 12):3581-4.
18. Nakahara K, Hataya T, Uyeda I. Inosine 5’-triphosphate can dramatically increase the yield of NASBA products targeting GC-rich and intramolecular base-paired viroid RNA. Nucleic Acids Res. 1998;26(7):1854-6.
19. Eiras M, Rodrigues-Silva S, Sanches-Stuchi E, Penteado-Natividade Targon ML, Alves-Carvalho S. Viroides em citros. Trop Plant Pathol. 2009;34(5):275-96.
20. Sieburth PJ, Irey M, Garnsey SM, Owens RA. The use of RT-PCR in the Florida citrus viroid indexing program. In: Duran-Vila N, Milne RG, Da Graça JV, editors. Proceedings XV Conference of the Internacional Organization of Citrus Virologist (IOCV). Riverside, CA; 2002. pp. 230-9.
21. Ito T, Ieki H, Ozaki K, Iwanami T, Nakahara K, Hataya T, et al. Multiple citrus viroids in citrus from Japan and their ability to produce exocortis-like symptoms in citron. Phytopathology. 2002;92(5):542-7.
22. Cañizares M, Marcos J, Pallás V. Molecular characterization of an almond isolate of hop stunt viroid (HSVd) and conditions for eliminating spurious hybridization in its diagnostics in almond samples. Eur J Plant Pathol. 1999;105(6):553-8.
23. WenXing X, Ni H, QiuTing J, Farooq AB, ZeQiong W, YanSu S, et al. Probe binding to host proteins: A cause for false positive signals in viroid detection by tissue hybridization. Virus Res. 2009;145(1):26-30.
24. Gómez G, Pallás V. A long-distance translocatable phloem protein from cucumber forms a ribonucleoprotein complex in vivo with hop stunt viroid RNA. J Virol. 2004;78(18):10104-10.
25. Rodio ME, Delgado S, Flores R, Di Serio F. Variants of peach latent mosaic viroid inducing peach calico: Uneven distribution in infected plants and requirements of the insertion containing the pathogenicity determinant. J Gen Virol. 2006;87(Pt 1):231-40.
26. Umaña R. Diagnóstico de CBCVd (Cocadviroide) y CVd-VI (Apscaviroide) en plantaciones citrícolas del Uruguay mediante técnicas de detección basadas en hibridación molecular no isotópica. Tesis de Maestría en Biotecnología. Facultad de Ciencias, Universidad de la República, Montevideo, Uruguay; 2010.
]]>27. Semancik JS, Morris TJ, Weathers LG, Rordorf GF, Kearns DR. Physical properties of a minimal infectious RNA (viroid) associated with the exocortis disease. Virology. 1975;63(1):160-7.
28. Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162(1):156-9.
29. Cañizares MC, Marcos JF, Pallás V. Studies on the incidence of hop stunt viroid in apricot trees (Prunus armeniaca) by using an easy and short extraction method to analyze a large number of samples. Acta Hort. 1998;472(1):581-7.
30. International Potato Center (CIP). Preparation of 32P-labeled probes by RNA transcription. In: Salazar LF, Jayasinghe U, editors. Techniques in Plant Virology. Training Manual. Sections 3, 4, 5. Lima: International Potato Center (CIP); 1997.
31. Manchester KL. Value of A260/A280 ratios for measurement of purity of nucleic acids. Bio-Techniques. 1995;19(2):208-12.
]]>32. Pagliano G, Orlando L, Gravina A. Detección y caracterización del complejo de viroides de cítricos en Uruguay. Agrociencia. 1998;1(2):74-83.
33. Mohamed ME, Hashemian SMB, Dafalla G, Bové JM, Duran-Vila N. Occurrence and identification of citrus viroids from Sudan. J Plant Pathol. 2009;91(1):185-90.
34. Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. New York: Cold Spring Harbor Laboratory Press; 1989.
35. Semancik JS, Tsuruda D, Zaner LJ, Geelen LMC, Weathers LG. Exocortis Disease: Subcellular distribution of pathogenic (viroid) RNA. Virology. 1976;69(2):669-76.
36. Li SF, Onodera S, Sano T, Yoshida K, Wang GP, Shikata E. Gene diagnosis of viroids: Comparisons of return-PAGE and hybridization using DIG-labeled DNA and RNA probes for practical diagnosis of hop stunt, citrus exocortis and apple scar skin viroids in their natural host plants. Ann Phytopathol Soc Jpn. 1995;61(4):381-90.
]]>37. Wang X, Zhou C, Tang K, Lan J, Zhou Y, Li Z. Preliminary Studies on Species and Distribution of Citrus Viroids in China. Agric Sci China. 2008;7(9):1097-103.
38. Tuiskunen A, Leparc-Goffart I, Boubis L, Monteil V, Klingstrom J, Tolou HJ, et al. Self-priming of reverse transcriptase impairs strand-specific detection of dengue virus RNA. J Gen Virol. 2010;91(4):1019-27.
39. Rodríguez R, Ramos PL, Dorestes V, Velázquez K, Peral R, Fuentes A, et al. Establishment of a non-radioactive nucleic acid hybridization technique for Begomovirus detection. Biotecnol Apl. 2003;20(3):164-9.
]]> Received in October, 2012.
Rodolfo Umaña. Departamento de Biología Vegetal, Facultad de Agronomía, Universidad de la República, Garzón 780, Montevideo, Uruguay. E-mail: rumana@fagro.edu.uy.
]]>