Se describe en este trabajo la detección de Escherichia coli enterotoxigénica LT(+). Este método está basado en la amplificación de un fragmento de DNA de 400 pares de bases mediante la reacción en cadena de la polimerasa (PCR). Los oligonucleótidos fueron diseñados por los autores y los patrones característicos se observaron cuando las muestras fueron sometidas a una electroforesis en gel de agarosa al 2 %. La PCR tuvo resultados positivos con las cepa de colección Escherichia coli O:149 K: 88 (LT+) y con 20 cepas aisladas de pacientes con diarreas agudas. Se observaron resultados negativos con Escherichia coli O:101 K:99 NM (ST+), Vibrio cholerae 01 y Acromons hydrophila.
Palabras clave: ESCHERICHIA COLI/aislamiento y purificación, REACCIÓN EN CADENA POR POLIMERASA.
Toxigenic Escherichia coli is a major cause of acute diarrhoea in children under five years old. Plasmid codify the genetic information for LT toxin.1 Laboratory test for LT toxin identification are laborious and time consuming, making somehow to reconize the sources of infection.2
The polymerase chain reaction (PCR) is a porwerful techique used to amplying specific segments from small amounts of genomic and plasmid DNA.3
We report the detection of Escherichia coli (LT+) producing character, this methods is based on amplying 400 base pair DNA fragment PCR with primers designed by the authors yielded a characteristic pattern when amplified samples were subjected to electrophoresis on 2 % agarose gel. Suspension of 106 bacterial cells from each strains heated at 95 oC foe 10 min and used in the PCR amplification was done in 100 mL reaction mixture containing 10 mM Tris HCL pH 8,3; 50 mM KCL; 2,5 mM MgCl; 0,4 pmoles primers: 10 mL target samples; 2,5 m Taq polymerase. The thirty cicles PCR consisted of a rapid three steps process of denaturation (95 oC) for 60s, anneling (55 oC) for 90s and extention (72 oC) for 45s using a GENE Controller (Pharmacia).4 Two primers employed to amplify the b400 bp segment which codifies the subnit B of E. coli (LT) with HPLC purification.
The oligonucleotide sequence are:
oligo P1: 5´d (CCCAATTCCAATTCAAATTATCA) 3'
OLIGO p2: 5' (CCC ATCCTACCATTACACACATCTC) 3'
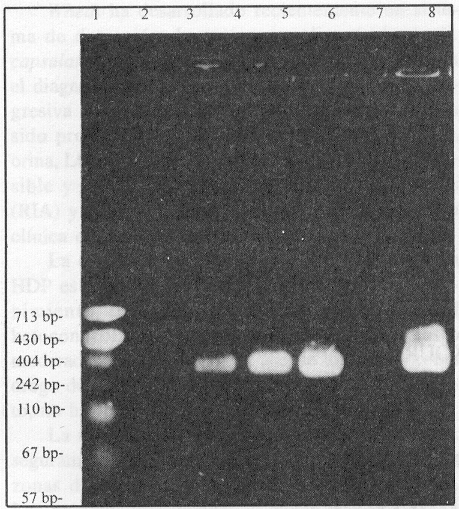
]]> The PCR amplification was successful when collection strains Escherichia coli O: 149 K: 88 (LT) as wellas twenty strains of patients with diarrhea were (figure), but not Aeromonas hydrophila, and Vibrio cholerae 01.FIGURE . PCR to identify toxigenic Escherichia coli (LT+). Line 1: DNA size marker of base pair 57,67,110,247,157,190, (Hpa II digest of Bs+). Line 2: empty lane. Line 3: amplified segment of toxigenic E. coli O:149 K:88 (LT+). Lines 4,5,6: amplified segment of toxigenic E. coli (LT+) isolated from patients with diarrhea diseases.
The specifity of the PCR was corroborated through restriction enzymes analysis5 for this purpose we used AccI and Hind II enzymes (CIGB) cleaved DNA at 90 bp and 180 bp.
In this paper it is described the detection enteroxigenic Escherichia coli LT (+). This method is based on the amplification of a DNA fragment of 400 pairs of bases by polymerase chain reaction (PRC). The oligonuclotides were designed by the authors and the characteristic patterns were observed when the samples were submitted to an electrophoresis in an Agarose gel at 2 %. The PCR had positive results with the strains of Escherichia coli 0:149 K; 88 (LT+) collection and with 20 strains isolated from patients with acute diarrhea. Negative results were found in Escherichia coli 0:101 K:99 NM (ST+), Vibrio cholerae 01 and Aeromonas hydrophilia.
Key words: ESCHERICHIA COLI/isolation and purification; POLYMERASE CHAIN REACTION.
Recibido: 10 de enero de 1996. Aprobado: 18 de mayo de 1996.
Dra. Margarita Ramírez. Instituto de Medicina Tropical "Pedro Kourí". Apartado 601, Marianao 13, Ciudad de La Habana, Cuba.
1 Especialista de I Grado en Microbiología. Instituto de Medicina Tropical "Pedro Kourí" (IPK).
2 Especialista de II Grado en Microbiología. Investigador Auxiliar. IPK
3 Dr. en Ciencias Biológicas. Facultad de Farmacia. Universidad de Barcelona.
4 Licenciada en Ciencias Biológicas. Investigadora Auxiliar. IPK.
5 Especialista de I Grado en Microbiología. Investigador Auxiliar. IPK.
6 Técnica en Procesos Biológicos. IPK.