Istituto Zooprofilattico Sperimentale Della Sicilia A. Mirri
Dr. Maria Vitale,1 Dr. Fabrizio Vitale,2 Dr. Vincenzo Di Marco,3 Dr. Vittoria Currò,2 Dr. Gesualdo Vesco4 and Dr. Santo Caracappa2
We set a method targeting 16 rRNA gene consisting in a single polymerase chain reaction of 40 cycles which is specific for pathogenic leptospira. Negative polymerase chain reaction results were observed with nonpathogenic Leptospira (serovar patoc) and other bacteria species. By this method a survey on a population of autochthon swine herds had been conducted in
Key words: Animal Leptospirosis, PCR, MAT, Sicilian black swine.
Leptospirosis, caused by the spirochete Leptospira, is considered an important reemerging infectious disease worldwide. Leptospirosis is a zoonosis of ubiquitous distribution, caused by infection with pathogenic Leptospira species. Leptospira have the ability to survive in a wide range of environmental reservoirs, including mammalian hosts.1,2 In some regions of the world human leptospirosis outbreaks are an important health problem. In Salvador, Brazil, more than 300 cases of human leptospirosis are identified each year during the rainy season, and 15% of them die.3 In Italy some human leptospirosis cases are reported only in the northern regions mainly connected with contaminated water.4 A serological survey by microscopic agglutination test (MAT) on sera samples from northern and central Italy of several animal species showed sero-positive percentages going from 12.13 % in ovine, 9.46 % in swine to 0.48 % in cattle.5 A single limited serological survey had been conducted until now in Sicily on animal leptospirosis (Vesco G et al. Unpublished results). To obtain more data on animal leptospirosis in
Isolation has been made from urine samples immediately put on EMJH containing 5-fluoroacil at 100 micrograms/ml; kidney samples were homogenized in the same medium and then incubated as described in OIE manual.8
Molecular analysis: DNA was extracted from kidney and liver samples using Fast DNA Kit from Q BIO GENE and Rybolyser Homogenization according to manufactures instructions.
DNA from urine samples was extracted using 2 aliquots of 200 microliters of urine with Qiagen columns following the manufacture's protocol without any further treatment; at the end we put together the 2 aliquots and perform PCR on 16S rRNA gene. PCR analysis had been performed using 16S rRNA gene as PCR target (primers: LEPTO E1 GGGAAAAATAAGCAGCGATGTG and LEPTO E2-ATTCCACTCCATGTCAAGCC amplicon 571 bp).9
]]> The following program in the Applied Biosystem 9 700 thermal cycler was used: 94 °C for 5 minutes 1 cycle followed by 40 cycles at 1 minute at 94 °C, 1 minute at 60 °C, 1 minute at 72 °C. The final extension was at 72 °C for 5 minutes.DNA from the bacterial strains was extracted by Qiagen column affinity column.
The amplified fragments were visualised on 2 % agarose gel.
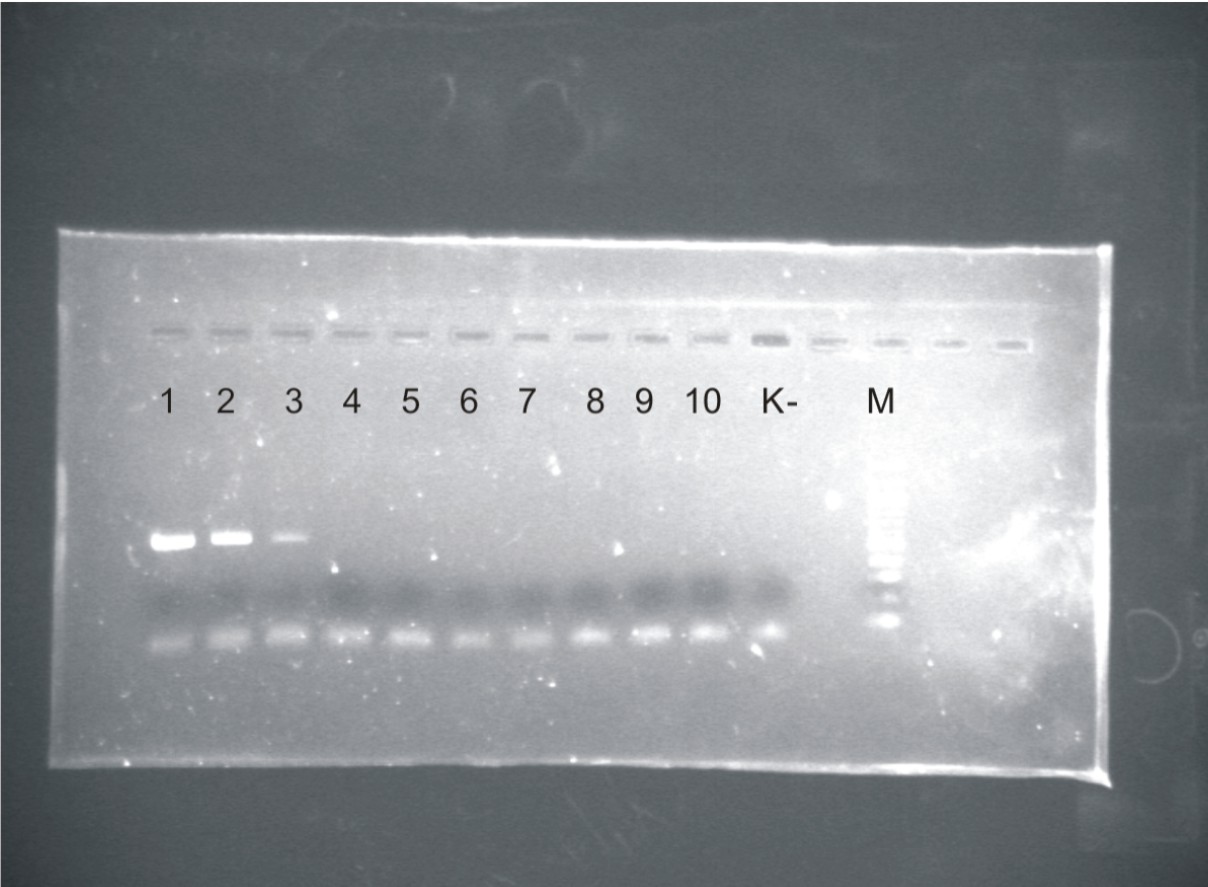
PCR method targeting 16S rRNA gene on DNA extracted from tissue and urine samples resulted in a high specific analytic method as shown in figure; pathogen Leptospira spp. were detected while several other bacteria species were negative.
Fig. PCR results on 1. L.pomoma; 2. L.grippothiphosa; 3. L.ballum; 4. E.coli; 5. B. melitensis; 6. S. aureus; 7. L. patoc; 8. S. abortus ovis; 9. M. bovis; 10. L. infantum; K-no DNA tamplate; M-100bp ladder
We performed analysis on different samples (urine, tissues) from black swine of Nebrodi to look for a prevalence of leptospirosis. In this population of animals we found PCR positive animals in a high percentage up to 40 % (Table 1).
Table 1. PCR and isolation data on samples from Black swine of Nebrodi.
| Kidney | ]]> PCR + | 30/90 33 % |
| Isolation + | 8/90 8,8 % | |
| Urine | PCR + | 28/ 70 40% |
| Isolation + | 15/70 21 % | |
| Liver | ]]> PCR + | 5/13 38% |
| Isolation + | 0/13 |
The animals were coming from a restricted area in
Survey on tissue and urine samples in bovine and ovine herds coming from the same areas in
Table 2. PCR and isolation data from bovine and ovine samples in
| Bovine | Ovine | ||
| ]]> Kidney | PCR + | 3/60 (5 %) | 3/45 (6,6 %) |
| Isolation + | 0/60 | 0/45 | |
| Urine | PCR + | 5/60 (8,3 %) | ]]> 3/60 (5 %) |
| Isolation + | 0/60 | 0/60 | |
The PCR targeting 16S rRNA gene can be a useful tool for the rapid diagnosis of leptospirosis detecting specifically many pathogenic leptospira. We found that the method is quite specific and analysis on DNA extracted by tissue samples and urine gave a higher number of positive animals compared to the isolation procedures. A previous serological screening on 110 bovine herds in south-east
The authors thank to M. Piazza, M. Leonardi and A. Fascetto for their technical assistance.
The work was financed by a grant to M.Vitale Ricerca corrente 2001 from the Italian Ministry of Health.
Se fijó un método que apuntó al gen del rRNA 16S, que consistió en una sola reacción en cadena de la polimerasa de 40 ciclos que es específica para leptospiras patógenas. Los resultados negativos de la reacción en cadena de la polimerasa fueron observados en leptospiras no-patogénicas (serovar patoc) y en otras especies de bacterias. Un estudio en una población de manadas de cerdos autóctonos en Sicilia se realizó por este método, particularmente en muestras de riñón de animales sacrificados y en muestras de orina de animales vivos. El análisis demostró una prevalencia de leptospiras de 40 % en estos animales. En otra manada de bovinos y ovinos, en la misma provincia de Sicilia, se encontró una baja prevalencia.
Palabras clave: Leptospirosis animal, RCP, MAT, cerdos negros de Sicilia.
Recibido: 27 de diciembre de 2004. Aprobado: 10 de marzo de 2005.
Dr. Maria Vitale. Head of Laboratory Genetica dei Microorganismi. Area Biologia Molecolare. Istituto Zooprofilattico Sperimentale della Sicilia A.Mirri. Via Gino Marinuzzi 4, 90129
1 Doctor in Veterinary Medicine. Physicians Doctors. Molecular Biology Istituto Zooprofilattico Sperimentale della Sicilia A. Mirri (Izs, Sicilia)
2 Doctor in Veterinary Medicine. Izs, Sicilia A. Mirri Palermo, Italy
3 Doctor in Biological Sciences. Izs, Sicilia A. Mirri