Ultrastructural remodelling of e ndoplasmic reticulum-derived membranes and hepatitis C virus morphogenesis in human liver samples
Modificación ultraestructural de las membranas derivadas del retículo endoplasmático y morfogénesis del virus de la hepatitis C en muestras de hígado humano
Viviana Falcón Cama,I Nelson Acosta-Rivero,I Sirenia González Pozos,II Santiago Dueñas-Carrera,I Emilio Acosta Medina,III Juan Kourí FloresII
I Centro de Ingeniería Genética y Biotecnología. Habana, Cuba.
II CINVESTAV-IPN. México, DF, México. ]]>
III Center for Advanced Study in Cuba. Habana, Cuba.
ABSTRACT
Introduction: hepatitis C virus infection is considered as a leading cause of liver disease worldwide. Despite recent advances in understanding the hepatitis C virus life cycle using the highly replicative JFH1 strain in human hepatoma cells, little is known about hepatitis C virus morphogenesis. Low levels of hepatitis C virus assembly in this cell culture model as well as low levels and complexity of hepatitis C virus particles in infected humans and chimpanzees have hampered the study of hepatitis C virus morphogenesis in vivo.
Objetivo: to study the ultrastructural features and viral assembly events in hepatocytes from HCV hepatitis C virus-infected patients.
Methods: liver needle biopsies samples of patients with hepatitis C virus infection, specific antibodies against hepatitis C virus and transmission electron microscopy and immunoelectron microscopy analyses were used in this study.
Results: ultrastructural studies in liver biopsies from hepatitis C virus-infected patients revealed that hepatitis C virus infection was related with remodelling of endoplasmic reticulum-derived membranes and with a variety of cytoplasmic microenvironments in hepatocytes. Dilated endoplasmic reticulum and formation of various membrane vesicles are features that have been associated with the viral replication complex. Interestingly, hepatitis C virus-like particles and core-like particles budding and assembly were observed near convoluted electron-dense membranes and tubular structures. Particularly, hepatitis C virus structural proteins localize to the endoplasmic reticulum.
Conclusions: these events indicate that hepatitis C virus nucleocapsids and early virion assembly take place at endoplasmic reticulum membranes that are associated with these cytoplasmic microenvironments in human hepatocytes.
RESUMEN
Introducción: la infección por el virus de la hepatitis C es una de las causas principales de la enfermedad del hígado a nivel mundial. La utilización de la cepa del virus de la hepatitis C, JFH1 en cultivo de células de hepatoma ha permitido el avance de la comprensión del ciclo de vida viral. No obstante, se conoce poco sobre la morfogénesis del virus de la hepatitis C. Las dificultades para detectar el ensamblaje viral en este modelo de cultivo celular, así como los bajos niveles y complejidad de las partículas del virus de la hepatitis C en pacientes y chimpancés infectados limitan el estudio de la morfogénesis viral.
Objetivo: estudiar las características ultraestructurales y los eventos de ensamblaje viral en hepatocitos de pacientes infectados con el virus de la hepatitis C.
Métodos: se utilizaron muestras de biopsias de hígado de pacientes infectados, anticuerpos específicos para el virus de la hepatitis C y técnicas de microscopía e inmunomicroscopía electrónica.
Resultados: la infección por el virus de la hepatitis C se relacionó con una modificación de las membranas derivadas del retículo endoplasmático y con diferentes microambientes citoplasmáticos en los hepatocitos de individuos infectados. La dilatación del retículo endoplasmático y la formación de diferentes vesículas de membrana son características que se asocian con los complejos de replicación viral. Resulta interesante destacar la detección del ensamblaje de partículas semejantes a la cápsida y al virus de la hepatitis C cerca de complejos de membrana con alta densidad electrónica y estructuras tubulares. Las proteínas estructurales del virus de la hepatitis C se detectaron en el retículo endoplasmático .
Conclusiones: estos eventos sugieren que el proceso temprano de ensamblaje de las nucleocápsidas y del virión ocurre en las membranas del retículo endoplasmático que se asocian con estos microambientes citoplasmáticos en los hepatocitos humanos.
Palabras clave: VHC; morfogénesis; replicación viral; biopsias de hígado; ensamblaje viral.
]]>
INTRODUCTION
Hepatitis C virus (HCV) infection is a major health problem (130-170 million people are estimated to be chronically infected) and is considered as a leading cause of liver disease worldwide.1 Most of infected persons develop chronic disease (70-80 %) that can progress to cirrhosis, hepatocellular carcinoma (HCC) and liver failure.1 In addition, it is predicted that the prevalence of HCV-related cirrhosis and HCC will considerably increase in the next decade.2 HCV was first identified in 1989 using molecular biology techniques from sera of experimentally infected chimpanzees.3 HCV is an enveloped positive-strand RNA virus classified as the type member of the Hepacivirus Genus in the family Flaviviridae.4 The HCV genome is a 9.6 kb single-stranded positive sense RNA that contains a single open reading frame (ORF) encoding a polyprotein of 3 010-3 030 amino acids (aa) residues.5 The ORF is flanked by 5´- and 3´-terminal no translated regions (NTRs) important for viral RNA translation and replication. The 5´-NTR contains an internal ribosome entry site (IRES) allowing translation of the RNA genome in the absence of a cap structure.6 Upon its synthesis in the endoplasmic reticulum (ER), the HCV polyprotein is co- and post-translationally processed by cellular and viral proteases into at least 10 mature cleavage products, including the structural (core, E1 and E2) and the non-structural (NS) proteins (p7, NS2, NS3, NS4A, NS4B, NS5A and NS5B). Core, E1, E2, p7 and NS2 are primarily involved in HCV assembly while NS3 to NS5B are primarily involved in viral RNA replication.5 HCV isolates show substantial genetic diversity and have been grouped into seven genotypes and several subtypes. HCV subtypes are epidemiologically distinct, with differences in risk group targeting and geographical distributions. Globally, genotypes 1 and 3 are the most common (46 % and 22 %, respectively).4
A major advance in HCV research occurred in 2005 with the discovery that the highly replicative HCV JFH1 strain (genotype 2a) was capable of producing infectious virions from selected hepatoma cell lines in culture.7-9 Ultrastructural studies from HCV cell culture systems disclosed that HCV induces a microenvironment, called the "membranous web" (MW), promoting viral replication.10 However, JFH1 is an unusual HCV strain allowing few assembly events per infected cell (in Huh7-derived hepatoma cells), thus hampering a better knowledge of the HCV assembly, morphogenesis and virion structure.11,12 In addition, little is known about HCV morphogenesis in the liver of virus infected persons. So, the goal of this work was to study the ultrastructural features and viral assembly events in hepatocytes from HCV-infected patients.
METHODS
PATIENTS AND SAMPLES
Liver needle biopsies from patients analysed in this study have been previously illustrated.13,14 Briefly, four patients with HCV infection hospitalized for hepatitis in the Institute of Gastroenterology, Havana, Cuba, were recruited after informed consent in writing was obtained (table). Liver needle biopsy samples were taken at the time of routine diagnostic biopsy from all patients. They were selected based upon they were serologically positive to third-generation HCV enzyme immunoassays (Tecnosuma International, Havana, Cuba) and that the anti-HCV positive sera were confirmed by Ortho HCV 2.0 ELISA (Ortho Diagnostic Systems, Rariton, NJ). They also showed positive detection of HCV RNA in the liver by in situ hybridization.14
]]> ANTIBODIES
Anti-core (CBSS-HepC.1) and anti-E1 (CBSS-HepC.2) mouse monoclonal antibodies (mAbs) have been described before.13,14
TRANSMISSION ELECTRON MICROSCOPY
Hepatic tissue samples were prepared for transmission electron microscopy as previously described.13,14 Briefly, hepatic tissue samples were fixed for 1 h at 4 _C in 1 % (v/v) glutaraldehyde and 4 % (v/v) paraformaldehyde, rinsed in 0.1M sodium cacodylate (pH 7.4), post-fixed for 1 h at 4_C in 1 % OsO4, and dehydrated in increasing concentrations of ethanol. The embedding was done, as previously described with minor modifications.13 In brief, ultrathin sections (400-500_AA) made with an ultramicrotome (NOVA, LKB) were placed on 400 mesh grids, stained with saturated uranyl acetate and lead citrate, and examined with a JEOL/JEM 2000 transmission electron microscope (JEOL, Japan).
Samples of hepatic tissue were fixed with 4 % (v/v) paraformaldehyde containing 0.2 % (v/v) glutaraldehyde in 0.1 M phosphate buffer (pH 7.3) at 4 _C for 3 h and washed with 0.1 M phosphate buffer, pH 7.3. Fixed cells were dehydrated as described above, embedded in Lowicryl, and polymerized by exposure to ultraviolet light at room temperature (RT) for 72 h. Ultrathin sections of liver biopsies were incubated with either anti-core (CBSSHepC.1) or anti-E1 (CBSS-HepC.2) mAbs in phosphate buffer, for 45 min at RT. The sections were rinsed three times for 30 min at RT with 0.1 % bovine serum albumin in phosphate-buffered saline, pH 7.3 (BSA-PBS), and incubated for 1 h at RT with gold-labeled (15 nm) anti-mouse IgG (Amersham, UK) diluted 1:100 in BSA-PBS. As control the primary antibody was substituted by normal mouse serum. All sections were stained and analysed with a transmission electron microscope, as mentioned above.
RESULTS
Ultrastructural studies in liver biopsies from HCV-infected patients revealed that HCV infection was related with remodelling of ER-derived membranes and with a variety of cytoplasmic microenvironments in hepatocytes (Fig. 1, 2 and 3). Ultrastructural features included dilated ER, formation of various membrane vesicles, convoluted electron-dense membranes and HCV-related tubular structures (Fig. 1 and 2). Particularly, HCV structural proteins localized to the ER membranes (Fig. 1, B and 2, B).
]]>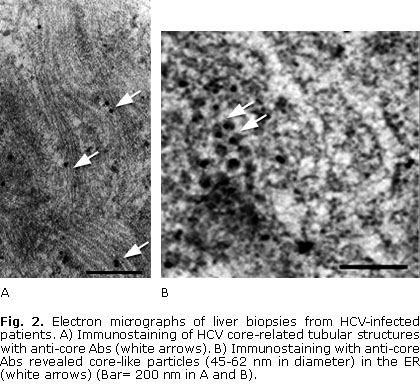
In addition, electron-dense ER membranes (that were immunolabelled with anti-E1 antibodies) displaying adhesion through their cytosolic sides, were observed in hepatocytes of HCV-infected patients (Fig. 1, B). Interestingly, both tubular structures and core-like particles were immunolabelled with anti-HCV core antibodies in hepatocytes (Fig. 2, A, B). Remarkably, budding of HCV-like particles suggesting viral assembly in ER membranes and near the tubular structures were shown in these liver samples (Fig. 2 and 3). Specially, enveloped particles with diameters between 68-80 nm, and electron-dense internal icosahedral-like capsids (diameters 25-35 nm) were observed (Fig. 3).
DISCUSSION
ULTRASTRUCTURAL REMODELLING OF ER-DERIVED MEMBRANES IN HEPATOCYTES DURING HCV INFECTION
To replicate its genome in in vitro systems, HCV alters ER-derived membranes to create in the cytoplasm a microenvironment, called the MW.15 The main constituents of the MW are single and predominantly double membrane vesicles (SMVs and DMVs).16,17 At later time points of HCV infection in cell culture, double-membrane tubules and multi-membrane vesicles (MMVs) appear, the latter presumably reflecting a stress induced host cell response.17 Results from this work and previous evidences suggest that HCV infection remodels ER-derived membranes in hepatocytes of HCV-infected patients to create in the cytoplasm a variety of microenvironments promoting viral replication and morphogenesis (Fig. 1, 2, 3).13 Notably, dilated ER and a variety of membrane vesicles are similar to those observed in vitro.
Several viral and host factors have been shown to be involved in MW formation. Biochemical and ultrastructural evidences indicate that both the MW and DMVs originate from the ER, with the outer membrane of DMVs connected by a neck to the ER membrane.13,17 In addition, NS4B has been shown to induce the formation of SMVs accumulations while NS5A induced DMVs.15,17 On the other hand, a number of host factors have been reported to contribute to the MW and DMVs formation, including nuclear pore complex (NPC) proteins, nuclear transport factors (NTFs), the autophagy pathway, and phosphatidylinositol 4-kinase IIIα (PI4KIIIα).18-20 Interestingly, NS5A and NS5B have been shown to interact with PI4KIIIα stimulating its activity. As a result, increased HCV replication and phosphatidylinositol 4-phosphate (PI4P) levels in the MW have been observed18 (Fig. 4).
HCV ASSEMBLY AND MORPHOGENESIS
The HCV core protein has been shown to induce the formation of convoluted ER membranes and HCV core-related tubular structures in cell culture, which have been associated with budding of nucleocapsids.21-23 HCV-related tubular structures have also been observed in hepatocytes from HCV-infected humans and chimpanzees.24,25 It is interesting to note that HCV-immunolabelled electron-dense ER membranes from hepatocytes of HCV-infected patients are similar to those found in the ER budding sites of HCV core particles in cells expressing the HCV structural proteins (Fig. 1, B).23 In addition, detection of core protein, budding core-like particles and HCV-like particles in ER membranes and near the tubular structures suggest they are related with early assembly of HCV (Fig. 2, 3 and 4). Notably, the observed enveloped particles (68-80 nm) with internal icosahedral-like capsids (diameters 25-35 nm) are similar to HCV core particles assembled in ER membranes of recombinant cells expressing the HCV structural proteins.26 Additional experimental evidences from human HCV-infected livers have shown that HCV core, envelope proteins and HCV RNA localize mainly to ER membranes in perinuclear areas of hepatocytes13,14,24 (Fig. 4). These experimental evidences indicate that ER membranes are the sites of HCV nucleocapsid and early virion assembly in hepatocytes from HCV-infected patients.
On the other way around, viral assembly has been related to HCV core interaction with lipid droplets (LDs) (formed either toward the cytosolic (cLDs) or luminal (luLDs) side of ER membranes) in JFH1 infected cells.27 Processing of HCV core on ER membranes by the SSP has been reported to be required for both core localization to cLDs and virus production.28 It is thought that HCV RNA replication and assembly occur in distinct membranous compartments (MW and ER, respectively). So, cLDs have been proposed to play a central role in the coordination of viral RNA synthesis and virion morphogenesis in the JFH1 cell culture model, by physically associating replication and assembly sites.29 In addition, LDs have been proposed as the site of HCV nucleocapsid assembly (11). Localization of HCV core and NS5A to cLDs and virus production is supported by diacylglycerol acyltransferase-1 (DGAT-1), an enzyme involved in triglyceride synthesis and luLDs maturation.30,31 In addition, heat shock cognate protein 70 (hsc70) and proteins associated with LDs, such as Rab18 and Tail-Interacting Protein 47 (TIP47), have been shown to colocalizes with HCV proteins promoting the interaction between sites of viral replication and LDs and the production of infectious virus particles.30,32,33 Moreover, viral double-stranded RNA has been found surrounding cLDs.34 Especially, NS5A has been suggested to be involved in the recruitment of the HCV replicase around cLDs and to deliver the HCV genome RNA to core protein35 (Fig. 4).
Nonetheless, it has been proposed that core protein must be retrieved from the surface of cLDs to move to the site of virus budding. Thus, a strong association of HCV core with cLDs has been related with reduced assembly of infectious particles.27 On the other hand, interaction of NS2 with E1-E2, p7, NS3 and NS5A has been shown to be essential for the migration of E1-E2 heterodimer, core protein and the NS3-NS4A enzyme complex at the virion assembly site (Fig. 4).36 In addition, rearrangement of E1E2 heterodimer complexes during the assembly of HCV particles has been suggested to yield a trimeric form of the E1E2 heterodimer.37 It is interesting to note that most studies suggesting a role of cLDs in virion assembly have used the JFH1 strain that produce scarce viral assembly events. Consequently, JFH1 chimeras carrying the structural region of different HCV strains have shown increased virus assembly and HCV core localization to ER membranes.21,38 Interestingly, 10 unusual amino acids in the JFH1 core sequence have been shown to affect viral assembly.21 By replacing these 10 residues in the JFH1 core with the most conserved residues, a significant increase in viral assembly and production was observed.21 Remarkably, this modified core protein mostly localized to the ER membranes showing increased ability to multimerize and to self-assemble into viral particles. Moreover, both E2 and NS5A colocalised with the modified JFH1 core on ER membranes suggesting that the early assembly events occurred at ER membranes distant to cLDs.21 This is similar to the ultrastructural findings in hepatocytes from human livers infected with HCV genotype 1b (Fig. 1, 2 and 3).13,14 Note that colocalization of HCV core with lipid droplets has been difficult to detect in hepatocytes of HCV-infected patients.13 Taken together, these experimental evidences indicate that HCV infection, both in cell culture and in infected persons, alters the lipid metabolism to induce substantial remodelling of ER membranes in hepatocytes (Fig. 4). Accordingly, HCV infection generates in the cell cytoplasm a variety of microenvironments promoting viral replication and morphogenesis. Ultrastructural features of the MW are similar in the HCV cell culture system and hepatocytes of infected individuals. However, differences are observed in microenviroments promoting HCV morphogenesis among cells infected with the JFH1 strain, those infected with chimeric HCV and hepatocytes from HCV-infected persons. Particularly, convoluted electron-dense membranes and HCV related tubular structures are related with HCV morphogenesis in hepatocytes of HCV-infected patients (Fig. 4).
CONCLUSIONS
Taken together, these experimental evidences indicate that HCV infection, both in cell culture systems and in humans, alters the lipid metabolism to induce substantial remodelling of ER membranes in hepatocytes. Accordingly, HCV generates a variety of microenvironments in the cell cytoplasm promoting viral replication and morphogenesis. Ultrastructural features of the MW are similar in the HCV cell culture system and hepatocytes of infected individuals. However, differences are observed in microenviroments promoting HCV morphogenesis among cells infected with the JFH1 strain, those infected with chimeric HCV and hepatocytes from HCV-infected persons. Particularly, convoluted electron-dense membranes and HCV related tubular structures are associated with HCV morphogenesis in hepatocytes of HCV-infected patients.
Conflicts of interest
The authors of the present paper hereby declare that there were no conflicts of interest. ]]>
Acknowledments
We would like to express our gratitude to Jesus Seone and Nilda Tamayo for their excellent technical assistance. The study was supported by the Center for Genetic Engineering and Biotechnology, Havana, Cuba.
REFERENCES
1. Mohd Hanafiah K, Groeger J, Flaxman AD, Wiersma ST. Global epidemiology of hepatitis C virus infection: new estimates of age-specific antibody to HCV seroprevalence. Hepatology. 2013;57:1333-42.
2. Kanwal F, Hoang T, Kramer JR, Asch SM, Goetz MB, Zeringue A, et al. Increasing prevalence of HCC and cirrhosis in patients with chronic hepatitis C virus infection. Gastroenterology. 2011;140:1182-8.
]]>3. Choo QL, Kuo G, Weiner AJ, Overby LR, Bradley DW, Houghton M. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science. 1989;244(4902):359-62.
4. Simmonds P. The origin of hepatitis C virus. Curr Top Microbiol Immunol. 2013;369:1-15.
5. Moradpour D, Penin F. Hepatitis C virus proteins: from structure to function. Curr Top Microbiol Immunol. 2013;369:113-42.
6. Niepmann M. Hepatitis C virus RNA translation. Curr Top Microbiol Immunol. 2013;369:143-66.
7. Lindenbach BD, Evans MJ, Syder AJ, Wolk B, Tellinghuisen TL, Liu CC, et al. Complete replication of hepatitis C virus in cell culture. Science. 2005;309:623-6.
]]>8. Wakita T, Pietschmann T, Kato T, Date T, Miyamoto M, Zhao Z, et al. Production of infectious hepatitis C virus in tissue culture from a cloned viral genome. Nat Med. 2005;11:791-6.
9. Zhong J, Gastaminza P, Cheng G, Kapadia S, Kato T, Burton DR, et al. Robust hepatitis C virus infection in vitro. Proc Natl Acad Sci, USA. 2005;102:9294-9.
10. Lohmann V. Hepatitis C Virus RNA Replication. Curr Top Microbiol Immunol. 2013;369:167-98.
11. Bartenschlager R, Penin F, Lohmann V, Andre P. Assembly of infectious hepatitis C virus particles. Trends Microbiol. 2011;19(2):95-103.
12. Lindenbach BD. Virion Assembly and Release. Curr Top Microbiol Immunol. 2013;369:199-218.
]]>13. Falcón V, Acosta-Rivero N, Chinea G, Gavilondo J, de la Rosa MC, Menendez I, et al. Ultrastructural evidences of HCV infection in hepatocytes of chronically HCV-infected patients. Biochem Biophys Res Commun. 2003;305(4):1085-90.
14. Falcón V, Acosta-Rivero N, Shibayama M, Luna-Munoz J, Miranda-Sanchez M, de la Rosa MC, et al. Evidences of hepatitis C virus replication in hepatocytes and peripheral blood monocuclear cells from patients negative for viral RNA in serum. Am J Infect Dis. 2005;1(1):34-42.
15. Gosert R, Egger D, Lohmann V, Bartenschlager R, Blum HE, Bienz K, et al. Identification of the hepatitis C virus RNA replication complex in Huh-7 cells harboring subgenomic replicons. J Virol. 2003;77:5487-92.
16. Ferraris P, Blanchard E, Roingeard P. Ultrastructural and biochemical analyses of hepatitis C virus-associated host cell membranes. J Gen Virol. 2010;91:2230-7.
17. Romero-Brey I, Merz A, Chiramel A, Lee JY, Chlanda P, Haselman U, et al. Three-dimensional architecture and biogenesis of membrane structures associated with hepatitis C virus replication. PLoS Pathog. 2012;8(12):e1003056.
]]>18. Berger KL, Kelly SM, Jordan TX, Tartell MA, Randall G. Hepatitis C virus stimulates the phosphatidylinositol 4-kinase III alpha-dependent phosphatidylinositol 4-phosphate production that is essential for its replication. J Virol. 2011;85:8870-83.
19. Dreux M, Garaigorta U, Boyd B, Decembre E, Chung J, Whitten-Bauer C, et al. Short range exosomal transfer of viral RNA from infected cells to plasmacytoid dendritic cells triggers innate immunity. Cell Host Microbe. 2012;12(4):558-70.
20. Neveu G, Barouch-Bentov R, Ziv-Av A, Gerber D, Jacob Y, Einav S. Identification and targeting of an interaction between a tyrosine motif within hepatitis C virus core protein and AP2M1 essential for viral assembly. PLOS Pathog. 2012;8:e1002845.
21. Etienne L, Blanchard E, Boyer A, Desvignes V, Gaillard J, Meunier JC, et al. The Replacement of 10 Non-Conserved Residues in the Core Protein of JFH-1 Hepatitis C Virus Improves Its Assembly and Secretion. PLoS One. 2015;10(9):e0137182.
22. Ait-Goughoulte M, Hourioux C, Patient R, Trassard S, Brand D, Roingeard P. Core protein cleavage by signal peptide peptidase is required for hepatitis C virus-like particle assembly. J Gen Virol. 2006;87:855-60.
]]>23. Blanchard E, Hourioux C, Brand D, Ait-Goughoulte M, Moreau A, Trassard S, et al. Hepatitis C viruslike particle budding: role of the core protein and importance of its Asp111. J Virol. 2003;77:10131-8.
24. De Vos R, Verslype C, Depla E, Fevery J, Van Damme B, Desmet V, et al. Ultrastructural visualization of hepatitis C virus components in human and primate liver biopsies. J Hepatol. 2002;37:370-9.
25. Shimizu JK, Weiner AJ, Rosenblatt J, Wong DC, Shapiro M, Popkin T, et al. Early events in hepatitis C virus infection of chimpanzees. Proc Natl Acad Sci, USA. 1990;87:6441-4.
26. Blanchard E, Brand D, Trassard S, Goudeau A, Roingeard P. Hepatitis C virus-like particle morphogenesis. J Virol. 2002;76:4073-9.
27. Shavinskaya A, Boulant S, Penin F, McLauchlan J, Bartenschlager R. The lipid droplet binding domain of hepatitis C virus core protein is a major determinant for efficient virus assembly. J Biol Chem. 2007;282:37158-69.
]]>28. Targett-Adams P, Hope G, Boulant S, McLauchlan J. Maturation of hepatitis C virus core protein by signal peptide peptidase is required for virus production. J Biol Chem. 2008;283:16850-9.
29. Bartenschlager R, Cosset F, Lohmann V. Hepatitis C virus replication cycle. J Hepatol. 2010;53:583-5.
30. Camus G, Herker E, Modi AA, Haas JT, Ramage HR, Farese RV, et al. Diacylglycerol acyltransferase-1 localizes hepatitis C virus NS5A protein to lipid droplets and enhances NS5A interaction with the viral capsid core. J Biol Chem. 2013;288:9915-23.
31. Herker E, Harris C, Hernandez C, Carpentier A, Kaehlcke K, Rosenberg AR, et al. Efficient hepatitis C virus particle formation requires diacylglycerol acyltransferase-1. Nat Med. 2010;16:1295-8.
32. Parent R, Qu X, Petit MA, Beretta L. The heat shock cognate protein 70 is associated with hepatitis C virus particles and modulates virus infectivity. Hepatology. 2009;49:1798-809.
]]>33. Vogt DA, Camus G, Herker E, Webster BR, Tsou CL, Greene WC, et al. Lipid droplet-binding protein TIP47 regulates hepatitis C Virus RNA replication through interaction with the viral NS5A protein. PLoS Pathog. 2013;9:e1003302.
34. Targett-Adams P, Boulant S, McLauchlan J. Visualization of double-stranded RNA in cells supporting hepatitis C virus RNA replication. J Virol. 2008;82:2182-95.
35. Moradpour D, Brass V, Penin F. Function follows form: the structure of the N-terminal domain of HCV NS5A. Hepatology. 2005;42:732-5.
36. Stapleford KA, Lindenbach BD. Hepatitis C Virus NS2 Coordinates Virus Particle Assembly through Physical Interactions with the E1-E2 Glycoprotein and NS3-NS4A Enzyme Complexes. J Virol. 2011;85:1706-17.
37. Lehner R, Lian J, Quiroga AD. Lumenal lipid metabolism: implications for lipoprotein assembly. Arterioscler Thromb Vasc Biol. 2012;32:1087-93.
]]>38. Galli A, Scheel TKH, Prentoe JC, Mikkelsen LS, Gottwein JM, Bukh J. Analysis of hepatitis C virus core/NS5A protein co-localization using novel cell culture systems expressing core-NS2 and NS5A of genotypes 1-7. J Gen Virol. 2013;94:2221-35.
Recibido: 18 de enero de 2016.
Aceptado: 4 de abril de 2016.
]]>