RESEARCH
Varying expression of neurotrophic factors in rat bone marrow stromal cells according to number of culture passages
Variación de la expresión de factores neurotroficos en celulas del estroma de medula ósea en ratas según el numero de paso del cultivo
Rocío García1, Nancy Pavón2, Paula Vergara3, José Segovia3, Esteban Alberti1
1 Departamento de Neurobiología
2 Departamento de Neuroinmunoquímica Centro Internacional de Restauración Neurológica, CIREN Ave. 25 # 15805, entre 158 y 160, Cubanacán, Playa, CP 11 300, Ciudad de La Habana, Cuba ]]>
3 Centro de Investigación y de Estudios Avanzados del Instituto Politécnico Nacional Ave. Instituto Politécnico Nacional 2508 Col. San Pedro Zacatenco, CP 07360, México, DF
ABSTRACT
Bone marrow stromal cells (BMSC) are multipotent stem cells and are considered good candidates for cell restoration in injured brain tissue. The aim of this study is to learn if rat BMSC express brain-derived neurotrophic factor (BDNF) and glia-derived nerve factor (GDNF) and determine if this expression varies according to the number of culture passages. The presence of BDNF and GDNF mRNA was determined by RT-PCR and protein expression was evaluated by Western Blot. GDNF production in individual cells in the culture was analyzed by Immunocytochemistry. Our results indicated that rat BMSC produce BDNF, at least until passage number 12. Nevertheless, GDNF production only takes place at passages 7 and 12; In conclusion, the expression of BDNF and GDNF by BMSC varies according to their mature state; both neurotrophic factors only are present after passage 7 which should be taken into account for their utilization as a therapeutic option of neurodegenerative diseases.
Keywords: bone marrow stromal cells, immunocytochemistry, neurotrophic factors, RT-PCR, western blot.
RESUMEN
Las células estromales de la medula ósea (CEMO) son células madre multipotentes y se consideran un buen candidato para la restauración celular del tejido cerebral dañado. El objetivo de este trabajo consistió en demostrar si las CEMO expresan los factores neurotróficos derivados de cerebro y de línea de células gliales (BDNF y GDNF, respectivamente) y determinar si esta expresión varía en relación con el número de pases en cultivo. La presencia del ARNm para BDNF y GDNF se determinó mediante RT-PCR y la expresión de la proteína se evaluó por Western Blot. La producción de GDNF por las células individuales en cultivo se analizó mediante Inmunocitoquímica. Nuestros resultados demostraron que las CEMO de rata producen BDNF, al menos hasta el pase 12. Sin embargo, la producción de GDNF solo tuvo lugar en los pases 7 y 12. En conclusión, la expresión de BDNF y GDNF por las CEMO varía en relación con su estado de maduración; sólo a partir del pase 7 están presentes ambos factores lo que debe tenerse en cuenta para la utilización de estas como opción terapéutica en enfermedades neurodegenerativas.
Palabras clave: celulas del estroma de medula ósea, inmunocitoquímica, factores neurotroficos, RT-PCR, western blot.
]]>
INTRODUCTION
Bone marrow stromal cells (BMSC) are adult stem cells with characteristics that make them an attractive candidate in cellular therapy for neurological diseases (1). The therapeutic potential of BMSC for the treatment of cerebral ischemia (2, 3), traumatic brain injury (4), and Parkinson´s disease (5) has been evaluated. These studies demonstrated that implanting BMSC into the injured brain induced a therapeutic improvement in animal models. The authors have suggested that the benefits observed in grafted animals are associated with the capacity of BMSC to produce neurotrophic factors.
Neurotrophic factors promote neuronal survival and stimulate axonal growth (6). In the adult brain, the lack of these proteins can induce apoptotic neuronal death (7, 8). It has been described that the brainderived neurotrophic factor (BDNF) regulates neurotransmitter release and dendritic and axonal growth. Moreover, BDNF rescues injured GABAergic neurons in animal models of Huntington´s disease (9). On the other hand, the glial cell line-derived neurotrophic factor (GDNF) has the strongest trophic effect on dopaminergic neurons (10). Studies of Parkinson´s disease in animal models, have shown the ability of GDNF to rescue damaged dopaminergic neurons after a lesion with neurotoxins (11, 12).
The expression pattern of these proteins in BMSC has not been well characterized. Reports in the literature, regarding the expression of BDNF and GDNF in BMSC, are limited and contradictory. Yang et al. did not detect these proteins in the supernatant of BMSC cultures (13), while other studies have demonstrated the expression of BDNF (14, 15) and GDNF (16). This may be associated to the differential expression of these neurotrophic factors depending on culture passage.
Considering that the production of neurotrophic factors in BMSC, implanted in the brain, may play an important role in the restoration of damaged tissue, the evaluation of the capacity of these cells to express BDNF and GDNF at different growth stages is relevant. This information could be useful for selecting the best number of passages at which BMSC should be implanted in animal models for neurological diseases.
MATERIALS AND METHODS
Isolation, culture and expansion of rat bone marrow stromal cells
Samples of BMSC were isolated from adult Wistar male rats. Whole bone marrow was extruded from femurs using a-MEM culture medium (Gibco). Clumps of bone marrow were centrifuged and the supernatant was discarded to remove debris. Pelleted cells were resuspended in a-MEM containing 10% fetal bovine serum (FBS) (Hyclone), 2mM L-glutamine (Gibco) and 100U/mL of penicillin-100 mg/mL of streptomycin (Gibco) and seeded in 25-cm2 culture flasks. After 48 h, non-adherent cells were removed by replacing the medium. Culture medium was changed every 4 days. When the culture monolayer reached a confluence of about 90%, cells were detached with 0.25% trypsin and replated.
Total RNA and protein isolation
]]> Once BMSC at passages 2, 7, and 12 had reached a confluence of approximately 90%, cell culture media was suctioned off and the surface was washed twice with phosphate buffer (NaCl 8 g/L; KCl 0.2 g/L; Na2- HPO4 1.09 g/L; KH2PO4 0.26 g/L, pH 7.2). Total RNA and proteins from cellular homogenates were extracted using Trizol (Roche Diagnostics). Protein concentration was quantified by Bicinchoninic Acid method.Reverse Transcriptase-Polymerase Chain Reaction Analysis
Complementary DNA (cDNA) from each sample was synthesized from 5 mg of total RNA primed with 0.5 mg oligo(dT)15 (Invitrogen). The mixture was heated to 70 °C for 10 min. First-strand buffer, 0.1 M DTT, and 25 mM dNTPs were added and the mixed contents were incubated at 42 °C for 2 min. One microliter of M-MLV reverse transcriptase (Invitrogen) was added and the mixture was incubated at 42 °C for 50 min. The reaction was inactivated by heating at 75 °C for 15 min.
PCR reactions were carried out using 2 mL of cDNA mixed with 25 mM dNTPs, 50 pmoles of each specific primer, 5 mL of DMSO, and 1U Taq DNA polymerase (Invitrogen). Cycling conditions were: 94 °C for 3 min; 40 cycles of 94 °C for 1 min, annealing temperature for 1 min, 72 °C for 1 min; and 72 °C for 5 min. The sequence of each primer, the annealing temperatures, and the length of the amplified products are given in table 1. Water was used as the negative control instead of cDNA. b-actin was used as an endogenous control. A 1 kb DNA ladder nucleic acid marker (Invitrogen) was used. Electrophoresis of DNA products was carried out on agarose gels at 100 mV and visualized with ethidium bromide (Table 1).
Analysis of BDNF and GDNF expression by Western Blot in cellular homogenates
Aliquots corresponding to 50 mg of total proteins of each sample were placed on electrophoresis on 12% denaturing polyacrylamide gels and transferred onto PVDF membranes (Bio-Rad Laboratories). The nonspecific binding of antibodies was blocked with 5% non-fat dried milk. Immunoblotting was carried out with rabbit polyclonal antibodies to BDNF or GDNF (1:200) (Santa Cruz Biotechnology), followed by HRP-labeled goat anti-rabbit IgG (1:2000) (Zymed). Immunoreactions were visualized using the chemilluminescence detection kit (ECL; Perkin Elmer, LAS, Inc). Membranes were stripped and incubated with monoclonal mouse antibody against b-actin (1:300) (17) and HRP-labeled goat anti-mouse IgG (1:7000) (Zymed).
Evaluation of GDNF expression by Immunocytochemistry
BMSC were grown on glass coverslips, fixed with 4% paraformaldehyde and treated with 0.2% triton X-100. Non-specific binding was blocked with 0.5% IgG-free bovine serum albumin (BSA). Coverslips were incubated with rabbit polyclonal antibody against GDNF (1:100) (Santa Cruz Biotechnology), followed by incubation with biotinylated goat antirabbit IgG (Vector Laboratories), and developed with Fluorescein-Streptavidin (Vector Laboratories). Cells incubated with 0.2% triton X-100 instead of the primary antibody served as negative controls. Coverslips were mounted on clean glass slides using Vectashield. Immunoreactive cells were visualized on an Olympus BX51 fluorescence microscope, and images captured using the Image Pro Acquisition and Analysis software (UVP). Cells were counterstained with DAPI (4´, 6-diamino-2-phenilindol, Vector Laboratories) to reveal the nuclei.
RESULTS AND DISCUSSION
Isolation, culture and expansion of rat BMSCs
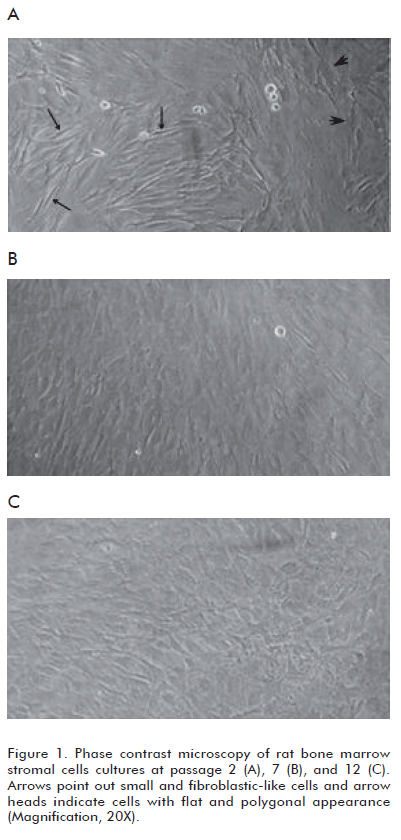
]]> We have observed that BMSC cultures are mainly composed of two morphologically distinct cellular populations and the prevalence of these populations differs among early and later passages. Cultures at passage 2 contained: a) cells with flat and polygonal appearance, and b) predominant small and fibroblastlike cells (Figure 1A). In contrast, BMSC cultures at passage 7 and 12 were more homogeneous, with a preponderance of fl at cells (Figures 1B and C). These results are in agreement with previous data obtained by other research groups (18-20).Observation of human BMSC cultures employing phase contrast microscopy has shown changes in the morphology of cells as a function of time in the culture and with the number of passages (18). Thus, fibroblast-like cells became fl at and polygonal, relatively mature, which predominate in cultures approaching senescence (19-21) (Figure 1).
Expression of BDNF in bone marrow stromal cells
Rat bone marrow stromal cells expressed BDNF as determined by RT-PCR (Figure 2A, upper panel) and Western blot (Figure 2B, upper panel), as early as passage 2. Moreover, these cells maintain the ability to produce this neurotrophic factor during the three culture passages evaluated.
Studies regarding the ability of BMSCs to synthesize neurotrophic factors show contradictory results. Nerve growth factor (NGF), BDNF, and GDNF production in mice BMSCs have been evaluated in fresh culture media by ELISA. This study revealed that murine BMSCs produce NGF but not BDNF or GDNF (13). Nevertheless, it has been shown by RT-PCR that BMSCs from mice, after passage 3, are able to express NGF and BDNF (14). In a similar study performed in human stromal cells the existence of BDNF mRNA was confirmed (15). Comparable results have been obtained in rat BMSC cultures at passage 5, where the presence of NGF and BDNF mRNA has been documented (22).
Here we demonstrated the expression of BDNF in rat BMSC and this result agrees with several reports (14, 15, 22). We also confirmed the expression at the three passages evaluated, which had not been reported previously.
Our results contrast with those reported by Yang et al. (13). These authors did not detect BDNF in the culture media from fresh murine BMSC (13). This disagreement could be due to two factors: first, the sample used; and second, the species analyzed. Yang et al. assessed the presence of BDNF in the culture supernatant from fresh BMSC at early passages; where protein concentration, if present, may be very low. In contrast, we evaluated the presence of the protein in cellular homogenates where there should be a higher quantity of BDNF. On the other hand, comparisons of BMSC from different strains of mice demonstrated that certain properties are strain specific (23). These features include doubling time, differentiation potential, optimal growth media requirements, propagation rates, and the presence of surface epitopes. Hence, there may be differences in the expression patterns of neurotrophic factors between rat and mouse BMSC.
We observed less BDNF mRNA and protein as the number of passages in the culture increased (Figure 2A and 2B, upper panels). It has been documented that changes in cellular morphology are associated with changes in patterns of gene expression (20, 24). Hence, cultures enriched for fibroblastic-like cells at early passages could have a greater potential to produce BDNF than cultures with a preponderance of large and mature cells.
BDNF is initially synthesized as a preproprotein. The pre-domain is cleaved off, yielding proproteins which can undergo further post-translational modifications (25). The proBDNF may be cleaved intracellulary to release mature BDNF, but when BDNF is over-expressed, the proform is modified extracellulary (26). PreproBDNF, proBDNF, and mature BDNF are detectable by Western Blot as bands averaging 42, 32, and 14 kDa respectively (27, 28).
]]> In our study, only the fragment corresponding to the preproprotein was labeled in all BMSC passages evaluated. The lack of proprotein and mature BDNF recognition in this study could due to the possible release of these molecules into the extracellular space and only the preproprotein is present in cellular homogenates (Figure 2).Expression of GDNF in bone marrow stromal cells
Rat bone marrow stromal cells failed to express GDNF at passage 2, as assessed by RT-PCR. Amplification products of this transcript were only detected at passage 7 and 12 (Figure 2A, middle panel). Protein expression by Western Blot confirmed these results, the fragment corresponding to GDNF was not detected in any cell culture at passage 2 (Figure 2B, middle panel).
Taking into account that RT-PCR allows the detection of very low levels of mRNA; our data indicates that cultures of BMSC at passage 2 do not express GDNF. However, the presence of GDNF mRNA was detected in BMSC cultures at passages 7 and 12. Our group was the first to report GDNF expression by BMSC, and only a few studies have been published since then (16).
GDNF expression in genetically unmodified BMSC has been evaluated in two studies. Yang et al. found that fresh murine BMSC did not secrete GDNF (13). More recently, Ye et al. assessed the production of GDNF in rat BMSC at passage 2; after 3, 7, and 10 days in the culture (29). This study revealed that BMSC were able to secrete GDNF and that the production of this neurotrophic factor gradually increased with time in the culture, with a maximum at 10 days. These authors considered that this increase was associated to a larger number of cells and greater growth rate (29).
Our results are consistent with those of Yang et al. (13), but they differ from the results of Ye et al. (29). This difference could be due to the cell culture medium used in both experiments. Ye et al. cultured cells in DMEM containing 20% FBS whereas we used a-MEM supplemented with 10% of FBS. Studies show that BMSC are very sensitive to soluble factors from the culture medium (30). Hence, discrepancies between these results and ours might be due to culture conditions.
Western Blot using an antibody against GDNF revealed the presence of two bands. The mature form of this protein has a molecular weight of 16 kDa and the glycosylated form of the mature GDNF has molecular masses of 20 or 23 kDa (31). However, GDNF preferentially exists as homodimer (8) which forms a complex with several molecules of heparin. This glycosylated homodimer has molecular weights ranging from 33 to 45 kDa (32). These immunoreactive bands, which were detected in this study, were broad and this is characteristic of glycosylated GDNF (31).
Relative intensity of bands corresponding to these molecular species varies when increasing the number passages. We assume that this is because BMSC cultures are formed by a diverse repertoire of distinct subpopulations, with different degrees of maturity, and therefore different patterns of gene expression.
To evaluate if BMSC at passage 2 did indeed secrete GDNF but at very low levels, an immunocytochemistry study was performed. This approach is very sensitive and makes it possible to study protein expression of different cellular populations in the same culture. Results agreed with those mentioned above. The production of GDNF in BMSC at passage 2 was not detected, whereas cells in passage 7 and 12 expressed this neurotrophic factor (Figure 3). It is possible that BMSC at different stages of growth, undergo molecular reorganization processes in which they acquire the ability to produce GDNF (Figure 3).
Stem cells could be an important source for cellular restoration in degenerative and traumatic neuronal diseases. Neuronal survival and functioning depend on the availability of sufficient amounts of growth factors (33). For this reason, the ability of BMSC to produce BDNF and GDNF supports the use of these cells as biological therapeutic vehicles for the treatment of neurological disorders. BDNF and GDNF production in BMSC can facilitate their differentiation into neural phenotypes (22). Moreover, these characteristics may help restore neural circuits, either through the differentiation of these cells into neurons per se or by trophic support. Taking into consideration that BMSC at passage 7 have the ability to produce BDNF and GNDF, both known to be relevant to therapy in a variety of neurodegenerative diseases, we consider that stromal cells should be collected at this passage for implanting in animal models of these disorders.
]]> ACKNOWLEDGMENTSPart of this study was supported by a United Nations University-BIOLAC fellowship.
REFERENCES
1. Mezey E. Bone marrow-derived stem cells in neurological diseases: stones or masons? Regen Med 2007;2:37-49.
2. Zhang C, Li Y, Chen J, Gao Q, Zacharek A, Kapke A, et al. Bone marrow stromal cells upregulate expression of bone morphogenetic proteins 2 and 4, gap junction protein connexin-43 and synaptophysin after stroke in rats. Neuroscience 2006; 141:687-95.
3. Kurozumi K, Nakamura K, Tamiya T, Kawano Y, Ishii K, Kobune M, et al. Mesenchymal stem cells that produce neurotrophic factors reduce ischemic damage in the rat middle cerebral artery occlusion model. Mol Ther 2005;11:96-104.
4. Brodhun M, Bauer R, Patt S. Potential stem cell therapy and application in neurotrauma. Exp Toxicol Pathol 2004;56:103-12.
5. Pavón N, Blanco L, Martínez L, Castillo L, Cuétara K, García R, et al. Trasplante de células estromales en el modelo de lesión por 6-OHDA. Rev Neurol 2004;39:326- 34.
6. Calissano P, Matrone C, Amadoro G. Nerve Growth Factor as a paradigm of neurotrophins related to Alzheimer’ s Disease. Dev Neurobiol 2010;70:372-83.
7. Mínguez A, Escamilla F. Terapia celular y otras estrategias neurorregenerativas en la enfermedad de Parkinson (II). Rev Neurol 2005;41:684-93.
8. Landreth GE. Growth Factors. In: Siegel GJ, Albers RW, Brady ST, Price DL, editors. Basic neurochemistry. Molecular, cellular and medical aspects. 7th ed. Boston, Mass: Elsevier Academic Press: 2006, p. 471-84.
9. Yoshii A, Constantine-Paton M. Postsynaptic BDNF-TrkB signaling in synapse maturation, plasticity, and disease. Dev Neurobiol 2010; 70:304-22.
10. Barroso-Chinea P, Cruz-Muros I, Aymerich M, Rodríguez-Díaz M, Afonso- Oramas D, Lanciego J, et al. Striatal expression of GDNF and differential vulnerability of midbrain dopaminergic cells. Eur J Neurosci 2005;21:1815-27.
11. Hurelbrink C, Barker R. The potential of GDNF as a treatment for Parkinson´s disease. Exp Neurol 2004;185:1-6.
12. Kordower J. In vivo gene delivery of glial cell line–derived neurotrophic factor for Parkinson’ s disease. Ann Neurol 2003;53:S120-34.
13. Yang K, Yuan X, Sun J, Long Y, Lai EC. Bone marrow stromal cell as potential cellular therapy for treatment of Parkinson´s disease. Mov Disord 2002;17 Suppl. 5:18.
14. Yamaguchi S, Kuroda S, Kobayashi H, Shichinohe H, Yano S, Hida K, et al. The effects of neuronal induction on gene expression profile in bone marrow stromal cells (BMSC)-a preliminary study using microarray analysis. Brain Res 2006;1087:15-27.
15. Zhang J, Li Y, Chen J, Cui Y, Lu M, Elias SB, et al. Human bone marrow stromal cell treatment improves neurological functional recovery in EAE mice. Exp Neurol 2005;195:16-26.
16. García R, Aguiar J, Alberti E, Cuétara K, Pavón N. Bone marrow stromal cells produce nerve growth factor and glial cell line-derived neurotrophic factors. Biochem Biophys Res Commun 2004;316:753-4.
17. Garcia-Tovar CG, Perez A, Luna J. Biochemical and histochemical analysis of 71 kDa dystrophin isoform (Dp71f) in rat brain. Acta Histochem 2001;103:209-24.
18. Sekiya I, Larson B, Smith JR, Pochampally R, Cui JG, Prockop DJ. Expansion of human adult stem cells from bone marrow stroma: conditions that maximize the yields of early progenitors and evaluate their quality. Stem Cells 2002;20:530-41.
19. Prockop DJ, Sekiya I, Colter DC. Isolation and characterization of rapidly self-renewing stem cells from cultures of human marrow stromal cells. Cytotherapy 2001;3:393-6.
20. Colter D, Class R, DiGirolamo C, Prockop DJ. Rapid expansion of recycling stem cells in cultures of plastic-adherent cells from human bone marrow. Proc Natl Acad Sci USA 2000;97:3213-8.
21. Gregory C, Prockop D, Spees J. Nonhematopoietic bone marrow stem cells: Molecular control of expansion and differentiation. Exp Cell Res 2005;306(2):330-5.
22. Yaghoobi MMY, Mowla SJ. Differential gene expression pattern of neurotrophins and their receptors during neuronal differentiation of rat bone marrow stromal cells. Neurosci Lett 2006;397:149-54.
23. Peister A, Mellad JA, Larson BL, Hall BM, Gibson LF, Prockop DJ. Adult stem cells from bone marrow (MSCs) isolated from different strains of inbred mice vary in surface epitopes, rates of proliferation, and differentiation potential. Blood 2004;103:1662-8.
24. Ylostalo J, Bazhanov N, Prockop D. Reversible commitment to differentiation by human multipotent stromal cells (MSCs) in single-cell derived colonies. Exp Hematol 2008;36:1390-402.
25. Leßmann V, Brigadski T. Mechanisms, locations, and kinetics of synaptic BDNF secretion: An update. Neurosci Res 2009;65:11-22.
26. Barker PA. Whither proBDNF? Nat Neurosci 2009;12:105-6.
27. Spires TL, Grote HE, Varshney NK, Cordery PM, van Dellen A, Blakemore C, et al. Environmental enrichment rescues protein deficits in a mouse model of Huntington’ s Disease, indicating a possible disease mechanism. J Neurosci 2004; 24:2270-6.
28. Yang J, Siao C- J, Nagappan G, Marinic T, Jing D, McGrath K, et al. Neuronal release of proBDNF. Nat Neurosci 2009;12:113-5.
29. Ye M, Chen S, Wang X, Qi C, Lu G, Liang L, et al. Glial cell line-derived neurotrophic factor in bone marrow stromal cells of rat. Neuroreport 2005;16:581-4.
30. Gregory C, Ylostalo J, Prockop DJ. Adult bone marrow stem/progenitor cells (MSCs) are preconditioned by micro- environmental niches in culture: a two- stage hypothesis for regulation of MSC fate. Sci STKE 2005;2005(294):pe37.
31. Katoh-Semba R, Tsuzuki M, Yoshida A, Nakajima H, Kitajima C, Matsuda M. Distribution and immunohistochemical localization of GDNF protein in selected neural and non-neural tissues of rats during development and changes in unilateral 6-hydroxydopamine lesions. Neurosci Res 2009;59:277-87.
32. Moretto G, Walker DG, Lanteri P, Taioli F, Zaffagnini S, Xu RY, et al. Expression and regulation of glial-cell-line-derived neurotrophic factor (GDNF) mRNA in human astrocytes in vitro. Cell Tissue Res 1996;286:257-62.
33. Morís G, Vega J. Factores neurotróficos: Fundamentos para su aplicación clínica. Neurología 2003;18(1):18-28.
Received in October, 2009. ]]> Accepted for publication in August, 2010.
Rocío García, Departamento de Neurobiología Centro Internacional de Restauración Neurológica, CIREN Ave. 25 # 15805, entre 158 y 160, Cubanacán, Playa, CP 11 300, Ciudad de La Habana, Cuba. E-mail: rociogminiet@infomed.sld.cu
]]>