RESEARCH
Study and isolation of aerobic hydrocarbon-degrading bacteria from Cuban shorelines
Estudio y selección de bacterias aerobias degradadoras de hidrocarburos del petróleo aisladas de costas de Cuba
Yaima Barrios-San Martín, Silvia Acosta, Ayixon Sánchez, Antonio Toledo, Francisca González, Regla M García
]]> Laboratorio de Química y Biotecnología, Centro de Investigaciones del Petróleo, Ceinpet. Churruca #481, Cerro, CP 12 600, La Habana, Cuba.
ABSTRACT
The isolation of aerobic marine bacteria able to degrade hydrocarbons represents a promising alternative for the decontamination of oceanic and coastal environments. In the present work, twelve water and sediment samples from the Felton coastline in the Province of Holguín were collected and screened with Bushnell-Haas medium supplemented with light crude oil or with seawater supplemented with yeast extract and crude oil as a carbon source, obtaining twenty seven and six bacterial isolates respectively that were able to grow in these media. The obtained isolates were then subjected to selection in Bushnell-Haas medium supplemented with a heavy crude oil, selecting three strains able to degrade this hydrocarbon mixture within a period of seven days. Pure cultures of these strains were further used in crude oil biodegradability assays. Total petroleum hydrocarbon (TPH) degradation was evaluated through SARA analysis, employing gas chromatography with an FID detector and infrared spectroscopy to analyze the aliphatic and aromatic hydrocarbon fractions, respectively. All three stains removed more than 60% of the TPH and one of them showed the best degradation potential with figures above 65% for the entire hydrocarbon fraction, except resins. Two of the strains were also able to decrease C17:Pr and C18:Ph ratios to less than 50% in comparison to the abiotic control. Two of these strains were phenotypically identified as sp., and the remaining one as sp. The degradation potential exhibited by these new isolates warrants further studies on their possible application to decontaminate coastal environments affected by oil spills.
Keywords: marine bacteria, biodegradation of hydrocarbons, Felton.
El aislamiento de bacterias marinas aerobias como posibles degradadoras de hidrocarburos, es una variante muy prometedora para la descontaminación de mares y costas. Se recogieron doce muestras de agua y de agua con sedimentos de las costas de Felton (Holguín, Cuba). Se empleó el medio Bushnell-Haas con petróleo crudo ligero y, alternativamente, agua de mar suplementada con extracto de levadura y petróleo crudo, como fuentes de carbono. Se obtuvieron 33 cepas bacterianas, que se sometieron a un proceso de selección en el medio Bushnell-Haas suplementado con crudo pesado, de las que se seleccionaron tres porque degradaban los hidrocarburos en siete días. Su capacidad de degradación se evaluó a escala de laboratorio. La remoción de los hidrocarburos totales del petróleo (HTP) se determinó mediante el análisis de los saturados, los aromáticos, las resinas y los asfáltenos. Los análisis de las fracciones saturadas y las aromáticas fueron mediante cromatografía de gases con detector de ionización de llama y espectroscopia infrarroja, respectivamente. Las tres cepas seleccionadas removieron más del 60% de los HTP. Una de ellas mostró valores de remoción superiores al 65% en todas las fracciones, excepto en las resinas; mientras que dos de ellas disminuyeron las tasas de C17/pristano y C18/fitano a menos del 50% con respecto al control abiótico. Dos de las cepas se identificaron fenotípicamente como del género , y otra como del género . Las potencialidades biodegradadoras de estos microorganismos en la limpieza de costas marinas han generado nuevos estudios.
Palabras clave: bacterias marinas, biodegradación de hidrocarburos, Felton.
INTRODUCTION
Many natural microorganisms, both aerobic and anaerobic, are able to thrive on hydrocarbons as sole carbon source. Usually found at low concentrations in non-contaminated areas, their populations bloom in chronically contaminated environments [1]. Anaerobic microorganisms, however, are less versatile regarding their growth substrate and often display increased sensitivity toward heavy metals, hence playing a smaller role in biodegradation [2]. Most research on bioremediation technology has focused therefore on aerobic heterotrophic bacteria, due not only to the taxonomic diversity of hydrocarbon-degrading representatives from this group, but to their ability to use xenobiotic compounds as carbon source in pure cultures [3]. Taxonomic and metabolic variety notwithstanding, individual species seldom have the complete enzymatic toolset required to completely degrade the main organic compounds contaminating the ecosystem at any given time, and thus biodegradation usually proceeds through the concerted action of mixed populations or microbial consortia. Viewed as a whole, the latter possess the necessary genetic information to produce all the enzymes required to completely degrade complex hydrocarbon mixtures in damaged areas [4, 5].
Environmental biotechnology techniques have been readily used to remediate ecological disasters caused by large oil spills, such as those caused by the collision of tanker Exxon Valdez with Bligh Reef at Prince William Sound, spilling some 50 000 tons of crude oil in March 1989 [6], and the sinking of Prestige, which spilled 63 000 tons of crude and ended up affecting 1 900 km of French and Spanish coastlines in November 2002 [7].
Measures taken after the Exxon Valdez spill included the addition of nutrients in the form of fertilizers (Inipol EAP 22 and Custombler), which increased the rate of oil removal by three-fold [6]. Laboratory studies prompted by the Prestige spill that examined oil biodegradability in Sorrizo beach (Coruña, Spain) in order to use biostimulation and bioaugmentation techniques demonstrated that these methods could produce total petroleum hydrocarbon (TPH) degradation rates of 6 to 45% after a seven day period. In addition, enriched cultures containing indigenous microorganisms and crude oil from the Prestige as sole carbon and energy source degraded close to 90% of the TPH in two weeks [7]. These and other successes have prompted us to undertake an effort to isolate, select and identify aerobic bacteria from the coast of Felton (Holguín, Cuba), which has been impacted by oil spills, characterizing the hydrocarbon-degrading potential of the isolated strains.
MATERIALS AND METHODS
Sampling
The marine ecosystem at Felton is currently contaminated by oil spills (Figure 1). Twelve samples (four from water, eight from water with silt) were manually collected from this environment, bottled into sterile 1 L flasks, and stored at 4 °C. They arrived to the laboratory for further processing within the first 24 h after their collection.
]]> Strain isolationThe primary selection process employed two different discrimination protocols, denominated here protocol A (modified from [8]) and B (modified from [9]). In protocol A the samples were stirred at 350 rpm for 5 min, taking a 1 mL aliquot from each one afterwards and seeding it into 9 mL of Bushnell-Haas medium [10] with 1% Mesa 30 crude (water and silt at 0.5% v/v, total sulfur 0.90% m/m, density at 15 °C of 0.8735 g/cm3, density 30° API) as sole carbon source. Five replicates were used per sample. After static incubation for 21 days at 26 to 28 °C, 0.1 mL of the resulting cultures were spread with a Drigalsky spatula onto the surface of marine bacteria isolation medium plates (10.0 g glucose, 0.5 g peptone, 1.0 g yeast extract, 15.0 g agar, 750 mL seawater, 250 mL distilled water), using three replicates per sample, and the plates were incubated at 26 to 28 °C for seven days. In protocol B the samples were stirred at 350 rpm for 5 min, bottling 100 mL from each one afterwards into a sterile 250 mL threaded cap bottle, to which 6 mL of Mesa 30 crude and 0.1 g of yeast extract were then added. The bottles were incubated for seven days at 130 rpm and 30 °C, after which 10 mL from each resulting culture were inoculated into 100 mL of sterile seawater containing 6 mL of Mesa 30 crude and 0.1 g of yeast extract. These cultures were then further incubated for seven days under the same conditions, at the end of which the subculturing cycle was repeated once more. After concluding the three subculture cycles, 0.1 mL from each final subculture were spread with a Drigalsky spatula onto the surface of marine bacteria isolation plates, employing three replicates per sample. The plates were incubated at 26 to 28 °C for seven days.
In both protocols the marine bacteria isolation plates were periodically examined after 24 h of growth under a stereoscope, streaking onto separate plates all the colonies appearing during the seven day incubation period. Pure stocks were obtained from colonies isolated by streaking, verifying their homogeneity by Gram staining and through the examination of culture characteristics.
Phenotypic characterization of the strains
The strains were characterized phenotypically using previously described morphological, physiological and biochemical tests [11-14], using previously defined criteria to describe culture characteristics [13]. Morphological descriptions were based on bacterial shape, motility and pigmentation.
Responses to temperature were evaluated by growing the strains under examination, plated on marine bacteria isolation medium, at 4, 22, 30 and 46 °C for 7 days. Salt tolerance was estimated by seeding the test strains in marine bacteria isolation medium where seawater was replaced by distilled water and sodium chloride (NaCl) concentration was set at 0, 0.5, 1, 3, 5, 7 and 10%, growing the cultures at 22 °C for 48 h and then scoring them daily for bacterial growth. The cultures were discarded after 15 days.
Biochemical characterization of the strains comprised tests for the fermentation of glucose, lactose, sucrose and mannitol, as well as for the production of indole, gas and hydrogen sulfide. It also involved assays for catalase, oxidase, urease, gelatinase, amylase and hemolysin activity, and growth tests in Simmons’ citrate, Mc Conkey, CromoCen® CC and CromoCen® AGN base media (Center for Biopreparations, Biocen, Cuba). NaCl was added to specific growth media at a final concentration of 2%.
Selection
The strains were grouped according to matching morphological, physiological and biochemical criteria, preparing inocula from each selected strain by diluting a loopful from a young culture into 5 mL of sterile seawater to an approximate concentration of 1 x 106 cfu/mL, according to the McFarland scale. One-hundred microliters of these inocula were added to Bushnell-Haas agar medium containing 1% Mesa 30 crude as sole carbon source and tetrazolium triphenyl chloride (TTC) as growth indicator, incubating the resulting culture for 21 days at room temperature and environmental relative humidity. Three replicates were seeded per strain. The cultures were examined every 24 h, selecting all strains exhibiting vigorous growth within a seven day period for further study.
Hydrocarbon degradation capacity
]]> Inoculum preparationThe selected strains were transferred to tryptone-soy agar medium (TSA) plates supplemented with 2% NaCl and incubated at 30 °C for 24 h. Loopfuls from each of the resulting bacterial lawns were transferred to 15 mL of sterile seawater and homogenized by vortexing to a concentration of 106 cfu/mL, according to the McFarland scale; verifying the obtained bacterial concentration by plating serial dilutions using the method of Track Dilution. The obtained bacterial suspensions constituted the inocula.
Culture conditions
The inocula, diluted at 10% (v/v), were added onto 88% (v/v) Bushnel-Haas media containing 2% (v/v) Mesa 30 crude as sole carbon source, for a final volume of 150 mL. These cultures were incubated at 30 °C for 45 days, employing a non-inoculated culture medium supplemented with crude as abiotic control. Each strain was assayed in triplicate.
Hydrocarbon determinations
Hydrocarbon determinations were performed at day 45. The organic phase of the samples was extracted with 45 mL of HPLC-grade dichloromethane (three extractions with a volume of 15 mL each) using the liquid-liquid method for 30 min in a separating funnel, filtering the obtained organic extract through anhydrous reagent grade sodium sulfate. Saturated, aromatics, resins and asphaltenes (SARA) were analyzed following the ASTM D2007 and ASTM D2549 standards. Asphaltenes were precipitated with n-pentane.
The saturated hydrocarbon fraction, dissolved in n-hexane, was analyzed in a Philips PU 4400 gas chromatographer (Philips Scientific, United Kingdom) with a flame ionization detector (FID) and a capillary 30 m x 25 mm CP SIL 5CB column, using the following parameters: injector temperature, 300 °C; detector temperature, 320 °C; column gradient, 60 to 300 °C (6 °C/min).
Mono-aromatic and poly-aromatic hydrocarbon fractions were analyzed by FTIR spectroscopy in a Mattson spectrophotometer (PYE UNICAM, United Kingdom), using the NaCl window film method. All spectra were processed using the Omnic v5.2A software application.
RESULTS AND DISCUSSION
]]> Strain isolationSampling sites in Felton were selected so as to ensure that the isolated biota were in the presence of hydrocarbons in their natural environment. This probably explains the high number of hydrocarbon-degrading microorganisms identified through isolation protocols A and B, which totaled 33 strains. Culturing these strains in the presence of hydrocarbons (Mesa 30 crude) favored the expression of enzyme systems involved in their degradation and the preferential isolation of hydrocarbon-tolerant clones.
Phenotypic characterization of the isolated strains
Based on physiological and biochemical tests, 28 out of 33 strains (85%) could be successfully assigned to a genus. Finer taxonomic classification to the level of species was not pursued since, despite the existence of taxonomic identification schemes for microorganisms from marine ecosystems [12, 15], successfully identifying recently described genera and species of marine bacteria requires the application of molecular methods [16].
Two genera of Gram-positive bacilli were identified (Bacillus and Kurthia), as well as five genera of Gram-negative bacilli (Alcaligenes, Acinetobacter, Marinomonas, Pseudomonas and Azotobacter). Five strains belonging to the latter group could not be identified with the biochemical tests employed in this study.
Bacillus was the most abundant genus (14 out of 33 isolates, 42%). Many Bacillus species have been shown to be able to degrade hydrocarbons [17-19], both in terrestrial [17-20] and marine or aquatic environments [21-26]. B. thermoleovorans is able to degrade naphthalene [18], and there is a Bacillus sp. strain that degrades toluene [20, 26]. A microbiological survey of the western Cuban continental shelf found this species in both northern and southern locations [27]. BIOIL-FC, a bioproduct from the Center of Marine Bioproducts (Cebimar) of the Cuban Ministry of Science, Technology and the Environment, is based on a strain of Bacillus licheniformis [23].
Five of the 33 strains (15%) belonged to the Alcaligenes genus. Some species of this genus have been isolated from marine environments contaminated with hydrocarbons [5, 17, 28]. Colores et al. observed that the addition of surfactants to hydrocarbon-contaminated soils resulted in a compositional shift of the local bacterial population from Alcaligenes sp. to Rhodococcus and Nocardia [29]. It has been shown that strain WW1 of Alcaligenes denitrificans can degrade four-ring polyaromatic hydrocarbons [30]. These microorganisms can be found not only in soil and water samples, but in clinical specimens, occasionally.
Two of the isolated strains belonged to the Pseudomonas genus. Publications reporting the presence of this genus in hydrocarbon-contaminated ecosystems and describing its hydrocarbon-degrading abilities have appeared in the literature from the early nineties [17], although its numbers have increased as of late [19, 24, 25, 31, 32]. Pseudomonas sp. strains have been shown to degrade polycyclic aromatic hydrocarbons both in terrestrial [28, 33-35] and aquatic [36, 37] environments, and they are widely distributed among the most dissimilar ecosystems [17, 23, 25, 26, 28, 31-40]. Published reports indicate that they can also degrade phenanthrene in soils [38] as well as anthracene, phenol [41] and methyl methylbromide in marine environments [40].
Another two strains belonged to the genera Acinetobacter and Marinomonas, respectively. Recent studies on the degradation of hydrocarbons in water bodies have made reference to the former [26, 36, 37]; this genus has, in addition, been found in not only in terrestrial and marine environments, but in sewage as well. Representatives of the Marinomonas genus were isolated for the first time from sediments contaminated with polycyclic aromatic hydrocarbons in 2001 [42]. Some authors place some species of this genus into Alteromonas instead [17]. In any case, they live exclusively in coastal and oceanic marine environments, therefore requiring seawater for their culture in vitro and growing optimally at temperatures ranging from 20 to 40 °C.
Lastly, the genera Kurthia and Azotobacter were represented each by a single strain. We were unable to find previous mentions in the literature of the presence of these microorganisms in hydrocarbon-contaminated environments. Kurthia is a genus of Gram-positive bacilli described as environmental bacteria, whereas Azotobacter is a typical inhabitant of water bodies and soils. Optimum growth temperatures for both species fall within the 20-30 °C range.
]]> Strain selectionAll 33 isolated strains were grouped according to culture, morphological, physiological and biochemical parameters. At the end, 18 strains representing each unique combination of these parameters were chosen to be subjected to the selection process.
From day 3 onwards, strain F1FLC of Bacillus sp. exhibited vigorous growth. Visible growth was also detected for strains F10S1 and F9S of Alcaligenes sp. and Bacillus sp. respectively from day 4 onwards.
The bacteria growing in Varadero Venta heavy crudes have probably developed mechanisms to maintain membrane integrity in the presence of excessive hydrocarbon flow, increasing for instance membrane rigidity by decreasing its unsaturated fatty acids content and modulating the cis/trans conformational ratio of its phospholipids, and inducing the synthesis of membrane complexes that pump out hydrocarbons, in a manner homologous to that of many antibiotic-resistance bacteria [43, 44]. Hydrocarbons are lipophilic compounds that inhibit growth when present at high concentrations [45], inducing the bacterial stress response and a series of changes at the membrane, enzyme and protein levels [44, 46].
Characterization of the hydrocarbon degradation capacity of the selected strains
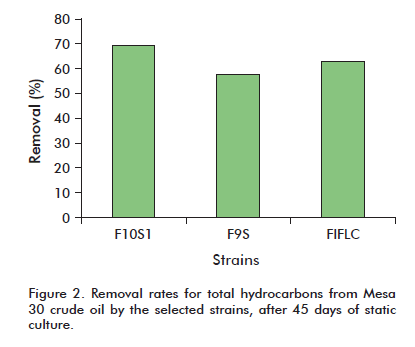
The ability to degrade hydrocarbons from crude oil (Mesa 30 crude) was determined after 45 days of static culture. Three of the strains under examination (F10S1, F9S and F1FLC) were able to remove over 50% of the starting hydrocarbon. Strain F10S1 degraded 69.26% of the crude (Figure 2).
The treatments (strains) exhibited statistically significant differences, both between them and in comparison with the control (p < 0.05). Although there were no significant inter-treatment differences regarding the concentration of aromatic hydrocarbon fraction I, the treatments did exhibit statistically significant differences when compared to the control (p < 0.05). No statistically significant differences were detected regarding the concentration of aromatic hydrocarbon fraction II or that of resins when comparing the treatments or even when these were compared to the control (p < 0.05). Asphaltene concentration, on the other hand, did exhibit statistically significant differences between treatments, as well as between these and the control (p < 0.05; Table 1).
Figure 3 shows the chromatographic profile of the saturated hydrocarbon fraction, in a signal intensity (pico-amperes) versus time chart for strains F9S, F10S1 and F1FLC. Each of the images shown in the figure contains the profiles of the abiotic control and that of the strain under examination, for a number of different carbon chain lengths. Those of strains F9S and F10S1 exhibit a higher decrease of maxima and of the non-resolved hydrocarbon background in comparison with strain F1FLC.
Chromatographic analysis was used to calculate C17/pristane and C18/phytane ratios for the different treatments and the control sample (Table 2).
]]> Monocyclic and polycyclic aromatic hydrocarbon fractions were analyzed by FTIR spectroscopy, as described in Materials and Methods. The transmittance (%) vs. wave number (cm-1) charts of strains F9S and F1FLC are shown in Figure 4; strain F10S1 exhibits a profile very similar to that of strain F9S (data not shown). Each chart depicts both the profile of the strain under examination and that of the standard sample. The concentration of associated hydroxyl groups (OH-, 3290-3300 cm-1) increased for all treatments, as did that of carboxylic groups (1680 cm-1).The present study has demonstrated that the bacterial strains selected are able to use hydrocarbons as sole carbon source when grown in pure cultures. Since hydrocarbon determinations were performed solely at the end of the study (day 45), no data are available to evaluate the biodegradation process in earlier time points. Based on our results, however, together with the existing literature, it is possible to make some inferences.
In general, biodegradation is expected to be more extensive in aliphatic hydrocarbons, which are far more amenable to this process than their aromatic counterparts [35, 47, 48]. The latter, in turn, are more susceptible to biodegradation than resins and asphaltenes [47, 48]. However, and despite the higher propensity of n-alkanes for oxidation [35, 49], no differences in biodegradation percentages were detected between saturated hydrocarbons, asphaltenes and monocyclic aromatic hydrocarbons after 45 days.
Aliphatic hydrocarbons decreased in comparison with those of the abiotic control regarding the non-resolved background (cycloalkanes, resins and asphaltenes). When C17/pristane and C18/phytane ratios of the strains under study were compared to those of the biological control, it was possible to detect a notable decrease for these values in strains F10S1 and F9S (50% approximately). This decrease in isoprenoids is a telltale sign of effective biodegradation; whereas strain F1FLC exhibits values above those of the abiotic control.
The spectra in figure 4 reveal the presence of aromatic compounds in the crudes treated with bacterial strains. No statistically significant differences were found for this parameter among the examined strains, although the increased proportion of carboxyl and hydroxyl groups demonstrates the presence of biological oxidation processes. The increased levels of phenols and phenoxides may be directly related to the accumulation of end compounds produced by the degradation of resins and asphaltenes.
All three strains under examination produced a notable decrease in the concentration of asphaltenes, compared to the control. This phenomenon was more pronounced for strains F10S and F9S, which did not exhibit statistically significant differences when compared in this regard, but did so when compared to strain F1FLC. The drop in asphaltene levels observed in the crude treated with strain F9S may have played a direct role in the increased amount of resins observed in this sample, although the removal rates of saturated and aromatic monocyclic hydrocarbons reached figures above 50%, and that of polycyclic aromatic hydrocarbons hovered around 40%. Strain F10S1 lowered the concentration of all fractions in comparison with the control. Samples treated with this strain exhibited the lowest concentration of polycyclic aromatic hydrocarbons, the second lowest concentration of asphaltenes and the third lowest of resins, and had removal rates over 65% for saturated and aromatic monocyclic hydrocarbons.
No alkanes with backbones shorter than 12 carbon atoms were detected in these samples. Narváez-Flores et al. made a similar observation while studying the hydrocarbon degradation capacity of bacteria isolated from marine sediments [44]. Several authors have pointed out that short chain aliphatic hydrocarbons usually volatilize during the first hours after a spill. Since their physico-chemical properties make them toxic compounds for the growth of most bacteria [44, 50], it is assumed that the degradation of aromatic compounds does not start until saturated hydrocarbons have been used up. However, direct experimental observation has revealed that low molecular weight aromatic compounds may start to be degraded much earlier, sooner, in fact, than many aliphatic molecules [51].
An analysis of these results leads us to suppose that linear chains up to 30 carbon atoms long and some low molecular weight aromatic compounds were degraded during the first 10 to 15 days. Had hydrocarbon composition been determined at that point, we would have most likely found practically identical levels of asphaltenes and resins in crudes treated with the strains under examination and in their controls, significantly decreased levels of saturated hydrocarbons in the former, and only a small drop in the concentration of aromatic hydrocarbons, since the bulk of their degradation takes place after 21 days. Ruberto et al., for example, were able to find statistically significant differences regarding the degradation of aromatic hydrocarbons between experimental treatments and their control only 20 days after the start of their bioaugmentation and biostimulation study [52]. The data published by Narváez-Florez et al., who failed to find statistically significant differences between hydrocarbon degradation rates in the abiotic control (3.6%) and the experimental treatment with bacteria (3.5%) [44] are also coherent with such a result.
When comparing the degradation of asphaltenes with that of saturated and aromatic hydrocarbons regarding the time point at which they first become detectable as well as their kinetics, it is useful to take into account the environmental characteristics of the habitat from which the microorganisms under examination have been isolated; that is, their environmental adaptations. In addition, it must be noted that synthesis of the enzyme complexes required to degrade the heavier fractions is not induced until lighter fractions have been exhausted, following the principle of maximum cellular economy (metabolic regulation). Regardless, Joseph et al. found that microorganisms isolated from the Cárdenas Bay in Matanzas (Cuba) degraded better heavy oils (such as the Varadero crude) than lighter ones (such as the Pina crude) [49]. These authors attributed such a phenomenon to adaptations of these microorganisms to the chronic contamination of their original habitats. Varadero crude is higher in asphaltenes, whereas the main constituents of Pina crude are saturated hydrocarbons with chain lengths smaller than 18 carbon atoms. Asphaltene degradation not only takes place through the classical, more efficient mechanisms of oxidation, but also through α- and ω-oxidation processes where intermediate products are bulkier, thereby lengthening the time required for biodegradation [23]. In addition, other authors have shown that the biotransformation of asphaltenes and resins leads to the accumulation of simpler saturated and aromatic derivates, increasing the concentration of these fractions [53, 54].
Bacterial metabolism is readjusted as culture ages and less complex substrates (saturated linear alkanes and low molecular weight aromatic compounds) are exhausted, shifting towards the synthesis of enzyme complexes geared towards the degradation of more complex molecules. These compounds did not mineralize completely, accumulating instead as intermediary metabolites consisting of linear chains and aromatic compounds. The host enzyme machinery must, therefore, have adapted to fluctuations in the levels of these different compounds.
]]>CONCLUSIONS
A total of 33 hydrocarbonoclastic strains were isolated from coastal environments at Felton (Holguín, Cuba). They were taxonomically identified as members of the Bacillus, Alcaligenes, Pseudomonas, Acinetobacter, Marinomonas, Kurthia and Azotobacter genera.
The strains Alcaligenes sp. F10S1; Bacillus sp. F9S and Bacillus sp. F1FLC removed 55 to 80% of the total hydrocarbon added to the culture after 45 days. Asphaltenes, monocyclic aromatic compounds and saturated hydrocarbons were the fractions undergoing degradation to the largest extent. Strains Alcaligenes sp. F10S1 and Bacillus sp. F9S exhibited the best results regarding the degradation of asphaltenes.
REFERENCES
]]>
1. Madigan M, Martinko J, Parker J. Brock, biología de los microorganismos. 10 ed. Revised. Madrid: Prentice Hall, Iberia; 2004.
2. Pollard SJT, Hrudey SE, Fedorak PM. Bioremediation of petroleum-and creosote-contaminated soils: a review of constraints. Waste Manage Res. 1994; 12(2):173-94.
3. Abalos A, Vinas M, Sabate J, Manresa MA, Solanas AM. Enhanced biodegradation of Casablanca crude oil by a microbial consortium in presence of a rhamnolipid produced by Pseudomonas aeruginosa AT10. Biodegradation. 2004;15(4):249-60.
4. Nápoles J. Ensayos de tratabilidad en suelos contaminados con petróleo [dissertation]. Santiago de Cuba: Universidad de Oriente; 2005.
5. Vinas M, Sabate J, Espuny MJ, Solanas AM. Bacterial community dynamics and polycyclic aromatic hydrocarbon degradation during bioremediation of heavily creosote-contaminated soil. Appl Environ Microbiol. 2005;71(11):7008-18.
]]>6. Swannell RP, Lee K, McDonagh M. Field evaluations of marine oil spill bioremediation. Microbiol Rev. 1996;60(2):342-65.
7. Fernandez-Alvarez P, Vila J, Garrido-Fernandez JM, Grifoll M, Lema JM. Trials of bioremediation on a beach affected by the heavy oil spill of the Prestige. J Hazard Mater. 2006;137(3):1523-31.
8. Bushnell LD, Haas HF. The Utilization of Certain Hydrocarbons by Microorganisms. J Bacteriol. 1941;41(5):653-73.
9. Baumann L, Baumann P, Mandel M, Allen RD. Taxonomy of aerobic marine eubacteria. J Bacteriol. 1972;110(1):402-29.
10. Rivera-Cruz MC, Ferrera-Cerrato R, Volke V, Rodríguez R, Fernández L. Adaptación y selección de microorganismos autóctonos en medios de cultivo enriquecidos con petróleo crudo. Terra Latinoam. 2002;20(4):423-34.
]]>11. Chablé de la Cruz G, García F, Madrigal EO. Evaluación de la respuesta de los microorganismos presentes en un derrame de hidrocarburos [Internet]. 2004 [cited 2009 Oct 29]. Available from http://www.biologia-en-internet.com
12. Oliver J. Taxonomic scheme for the identification of marine bacteria. Deep-sea Res. 1982;29(6):795-8.
13. Martínez J, Romay Z, Rojas T, Guerra G. Manual práctico de Microbiología. La Habana: Pueblo y Educación; 1985.
14. Pellón F, Orozco R, León J. Bacterias marinas con capacidad antimicrobiana aisladas de moluscos bivalvos en cultivos. Rev Per Biol. 2001 [cited 2010 Apr 14];8(2):[about 10 p.]. Available from: http://sisbib.unmsm.edu.pe/BVRevistas/biologia/v08_n2/bacte_marinas.htm
15. Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST. Bergey´s Manual of Determinative Bacteriology. 9 ed. Baltimore: Williams & Wilkins; 1994.
16. Gallardo AA, Risso S, Fajardo MA, Estevao S. Caracterización de polaciones microbianas presentes en la macroalga comestible Monostroma undulatum, Wittrock. Arch Latinoam Nutr. 54(3):337-45.
17. Leahy JG, Colwell RR. Microbial degradation of hydrocarbons in the environment. Microbiol Rev. 1990;54(3):305-15.
18. Annweiler E, Richnow HH, Antranikian G, Hebenbrock S, Garms C, Franke S, et al. Naphthalene degradation and incorporation of naphthalene-derived carbon into biomass by the thermophile Bacillus thermoleovorans. Appl Environ Microbiol. 2000;66(2):518-23.
19. Roling WF, Milner MG, Jones DM, Lee K, Daniel F, Swannell RJ, et al. Robust hydrocarbon degradation and dynamics of bacterial communities during nutrient-enhanced oil spill bioremediation. Appl Environ Microbiol. 2002;68(11):5537-48.
20. Ficker M, Krastel K, Orlicky S, Edwards E. Molecular characterization of a toluene-degrading methanogenic consortium. Appl Environ Microbiol. 1999; 65(12):5576-85.
21. Dojka MA, Hugenholtz P, Haack SK, Pace NR. Microbial diversity in a hydrocarbon- and chlorinated-solvent-contaminated aquifer undergoing intrinsic bioremediation. Appl Environ Microbiol 1998;64(10):3869-77.
22. Olivera NL, Commendatore MG, Moran AC, Esteves JL. Biosurfactant-enhanced degradation of residual hydrocarbons from slip bilge wastes. J Ind Microbiol Biot. 2000;25(2):70-3.
]]>23. Núñez R. Obtención, caracterización y aplicación de un bioproducto bacteriano para la biorremediación de derrames de hidrocarburos [dissertation], La Habana: Universidad de La Habana; 2004.
24. Ara-Rojas SL, Massol-Deyá A. Diversidad bacteriana en un biorreactor de lecho fluidificado durante el tratamiento de agua contaminada con nafta. Rev Argent Microbiol. 2007;39(4):243-51.
25. Kim JS, Crowley DE. Microbial diversity in natural asphalts of the Rancho La Brea Tar Pits. Appl Environ Microbiol. 2007;73(14):4579-91.
26. Winderl C, Anneser B, Griebler C, Meckenstock RU, Lueders T. Depth-resolved quantification of anaerobic toluene degraders and aquifer microbial community patterns in distinct redox zones of a tar oil contaminant plume. Appl Environ Microbiol. 2008;74(3):792-801.
27. Daane LL, Harjono I, Zylstra GJ, Haggblom MM. Isolation and characterization of polycyclic aromatic hydrocarbon-degrading bacteria associated with the rhizosphere of salt marsh plants. Appl Environ Microbiol. 2001;67(6):2683-91.
]]>28. Colores GM, Macur RE, Ward DM, Inskeep WP. Molecular analysis of surfactant-driven microbial population shifts in hydrocarbon-contaminated soil. Appl Environ Microbiol. 2000;66(7):2959-64.
29. Kanaly RA, Bartha R, Watanabe K, Harayama S. Rapid mineralization of benzo[a]pyrene by a microbial consortium growing on diesel fuel. Appl Environ Microbiol. 2000;66(10):4205-11.
30. Roling WF, Milner MG, Jones DM, Fratepietro F, Swannell RP, Daniel F, et al. Bacterial community dynamics and hydrocarbon degradation during a field-scale evaluation of bioremediation on a mudflat beach contaminated with buried oil. Appl Environ Microbiol. 2004;70(5):2603-13.
31. Bordenave S, Goni-Urriza MS, Caumette P, Duran R. Effects of heavy fuel oil on the bacterial community structure of a pristine microbial mat. Appl Environ Microbiol. 2007;73(19):6089-97.
32. Miravet ME. Estudio de bacterias heterótrofas en el Golfo de Batabanó [dissertation]. La Habana: Universidad de La Habana; 1996.
]]>33. Cerniglia CE. Biodegradation of Polycyclic Aromatic Hydrocarbons. Biodegradation. 1992;(2-3):351-68.
34. Dagher F, Deziel E, Lirette P, Paquette G, Bisaillon JG, Villemur R. Comparative study of five polycyclic aromatic hydrocarbon degrading bacterial strains isolated from contaminated soils. Can J Microbiol. 1997;43(4):368-77.
35. Whyte LG, Bourbonniere L, Greer CW. Biodegradation of petroleum hydrocarbons by psychrotrophic Pseudomonas strains possessing both alkane (alk) and naphthalene (nah) catabolic pathways. Appl Environ Microbiol 1997;63(9):3719-23.
36. Hilyard EJ, Jones-Meehan JM, Spargo BJ, Hill RT. Enrichment, isolation, and phylogenetic identification of polycyclic aromatic hydrocarbon-degrading bacteria from Elizabeth River sediments. Appl Environ Microbiol. 2008;74(4):1176-82.
37. Mukred AM, Hamid AA, Hamzah A, Wan Yusoff WM. Development of Three Bacteria Consortium for the Bioremediation of Crude Petroleum-oil in Contaminated Water. Online J Biol Sci. 2008; 8(4):73-9.
]]>38. Stringfellow WT, Aitken MD. Comparative physiology of phenanthrene degradation by two dissimilar pseudomonads isolated from a creosote-contaminated soil. Can J Microbiol. 1994;40(6):432-8.
39. Noordman W, Bruseau M, Jassen D. Effects of rhamnolipid biosurfactants on removal of phenanthrene from soil. Environ Sci Technol. 1998;32(12):1806-12.
40. Goodwin KD, Tokarczyk R, Stephens FC, Saltzman ES. Description of toluene inhibition of methyl bromide biodegradation in seawater and isolation of a marine toluene oxidizer that degrades methyl bromide. Appl Environ Microbiol. 2005;71(7):3495-503.
41. Neumann G, Teras R, Monson L, Kivisaar M, Schauer F, Heipieper HJ. Simultaneous degradation of atrazine and phenol by Pseudomonas sp. strain ADP: effects of toxicity and adaptation. Appl Environ Microbiol. 2004;70(4):1907-12.
42. Melcher RJ, Apitz SE, Hemmingsen BB. Impact of irradiation and polycyclic aromatic hydrocarbon spiking on microbial populations in marine sediment for future aging and biodegradability studies. Appl Environ Microbiol. 2002;68(6):2858-68.
]]>43. Ramos JL, Duque E, Rodriguez-Herva JJ, Godoy P, Haidour A, Reyes F, et al. Mechanisms for solvent tolerance in bacteria. J Biol Chem. 1997;272(7):3887-90.
44. Narváez-Florez S, Gómez ML, Martínez MM. Selección de bacterias con capacidad degradadora de hidrocarburos aisladas a partir de sedimentos del Caribe colombiano. Bol Invemar. 2008; 37(1):61-75.
45. LaGrega MD, Buckingham PL, Evans JC, Environmental Resources Management group. Hazardous waste management. 2nd ed. Boston: McGraw-Hill; 2001. p. 7-23.
46. Van Hamme JD, Singh A, Ward OP. Recent advances in petroleum microbiology. Microbiol Mol Biol Rev. 2003;67(4):503-49.
47. Song HG, Wang X, Bartha R. Bioremediation potential of terrestrial fuel spills. Appl Environ Microbiol. 1990;56(3): 652-6.
]]>48. León N, Infante C, Arias M, Marquez M, Gorrín A. Biodegradability of Venezuela crude oils. SPE International conference on health, safety, and environment in oil and gas exploration and production, 1998 June 7-10, Caracas, Venezuela.
49. Joseph IN, Capo MC, Bellota M, Ramos Y, Ramos I, Fuentes M. Aislamiento y selección de microorganismos degradadores de hidrocarburos en la plataforma cubana. Ciencias Biológicas. 1994;27:137-48.
50. Solano-Serena F, Marchal R, Casaregola S, Vasnier C, Lebeault JM, Vandecasteele JP. A Mycobacterium strain with extended capacities for degradation of gasoline hydrocarbons. Appl Environ Microbiol. 2000;66(6):2392-9.
51. Venosa A, Zhu X. Biodegradation of crude oil contaminating marine shorelines and freshwater wetlands. Spill Sci technol Bull. 2003;8:163-78.
52. Ruberto L, Vñazquez S, y Mac Cormack W. Effectiveness of the natural bacterial flora, bioestimulation and bioaumentation on the bioremediation of a hydrocarbon contaminated Antartic soil. Int Biodeter Biodegr. 2003;52(2):115-25.
53. Young LY, Cerniglia CE, editors. Microbial Transformation and degradation of toxic organic chemicals. Nueva York: Willy-Liss; 1995.
54. Araujo I, Gómez A, Barrera M, Angulo N, Morillo G, Cárdenas C, et al. Surfactantes biológicos en la biorremediación de aguas contaminadas con crudo liviano. Interciencia. 2008; 33(4):245-50.
Received in October, 2010.
Accepted for publication in January, 2012.
Yaima Barrios-San Martín. Laboratorio de Química y Biotecnología, Centro de Investigaciones del Petróleo, Ceinpet. Churruca #481, Cerro, CP 12 600, La Habana, Cuba. E-mail: yy@ceinpet.cupet.cu
]]>