Protocolo de purificación en dos etapas de fosfolipasas A2 a partir de la anémona marina Condylactis gigantea
A Two-Step Purification Procedure of Phospholipases A2 from the Sea Anemone Condylactis gigantea
MSc. Frenkel Guisado-BourzacI; MSc. Dolores Lázara Romero-Del-SolII, José Manuel Guisán-SeijasIII, Joaquín Díaz-BritoII; Dr.C. Andreimar Martins-SoaresIV, MSc. Alberto del Monte-MartínezII
IFacultad de Ciencias Naturales, Universidad de Oriente, Santiago de Cuba, Cuba, fguisado@uo.edu.cu, joaquin43cu@yahoo.es ]]>
IICentro de Estudio de Proteínas (CEP), Facultad de Biología, Universidad de La Habana, Ciudad Habana, Cuba, laz@fbio.uh.cu, jogoba@fbio.uh.cu, adelmonte@fbio.uh.cu
IIIInstituto de Catálisis y Petroleoquímica, CSIC, Campus UAM , Madrid, España, jmguisan@icp.csic.es
IVCentro de Estudos de Biomoléculas Aplicadas à Saúde, CEBio, Fundação Oswaldo Cruz, Fiocruz Rondônia e Departamento de Medicina, Universidade Federal de Rondônia, Porto Velho-RO, Brazil, andreimar@fiocruz.br
RESUMEN
El veneno de los celenterados marinos constituyen mezclas complejas de varios componentes, muchos de naturaleza proteica, y para los cuales se ha descrito la actividad fosfolipásica A2. El objetivo de la presente investigación fue purificar las fosfolipasas A2 (FLA2) procedentes de la anémona marina Condylactis gigantea a partir de un protocolo de dos etapas para su posterior caracterización. Se muestra la purificación de FLA2 en un soporte cromatográfico de afinidad con fosfatidilcolina de huevo inmovilizada covalentemente para la purificación de fosfolipasas A2, a los que se les comprobó cualitativamente la actividad fosfolipásica A2 por cromatografía en placa (TLC) utilizando sustrato marcado con fluorescencia. Dicho soporte cromatográfico permite que en un protocolo de purificación de solamente dos pasos se obtengan tres componentes proteicos de pesos moleculares entre 18000 y 14000, con al menos un componente que posee actividad fosfolipásica A2.
Palabras clave: cromatografía de afinidad, fosfolipasa A2, anémona marina, Condylactis gigantea, purificación de proteínas.
ABSTRACT
Marine coelenterate venom is composed of complex mixtures of several substances, mainly of proteins, for which phospholipase A2 activity has been described. This research study aims to test a two-step purification procedure for phospholipases A2 (PLA2) to filter enzymes for further characterization. PLA2 purification from the sea anemone Condylactis gigantean is conducted through a chromatographic affinity support MANA-Sepharose CL 4B with covalently immobilized phosphatidylcholine egg. The phospholipase A2 activity was corroborated by using qualitative TLC and a fluorogenic substrate. By means of the above-mentioned support, purification of three protein-based components can be carried out. These components are produced with molecular weights between 18000 and 14000, and at least one component possessing phospholipase A2 activity.
]]> Keywords: affinity, phospholipase A2, Condylactis gigantea, sea anemone, protein purification.
INTRODUCTION
Phospholipases A2 (PLA2) are esterolytic enzymes with high substrate and bond specificity for the hydrolysis of the sn-2 position of phosphoacylglycerols. These enzymes yield a lyso derivative (lysophospholipid) and a free-fatty acid molecule, which is able to hydrolyze both natural and synthetic phospholipids regardless of the nature of the fatty acids in the molecule. It also reveals maximum activity when the substrate is organized in large molecular aggregations such as micelles [1].
These enzymes are widely spread in nature [2, 3] and their functional versatility and usefulness have increasingly conditioned their use in Specialized and Applied Biochemistry as well as biotechnological, pharmaceutical, and general industries; such as food industry [4, 5], where a highly purified enzyme is not always needed.
In spite of their wide distribution in nature, most of these enzymes have been isolated from snakes, bees and scorpions venoms, as well as from pancreas tissues of mammals and membranes; for example, those of mitochondria from hepatic tissues [6].
Recent studies about marine coelenterates venom have demonstrated affinity to that from snakes, based on the pharmacological properties they possess, and the place where this enzymatic activity has been described for some species. The sea anemone Condylactis gigantean is a representative of this family.
Marine organisms have been less explored than mammals and microorganisms even though there are higher probabilities to find enzymes that have different specifics towards non-natural substrates in those environments. That is why the quest for phospholipase activity in marine invertebrates has been a focus in this field [7-10]. The Centro de Estudios de Proteinas in Havana has developed a two-step procedure in order to purify PLA2 from whole extracts of the sea anemone Condylactis gigantea.
]]> THEORETICAL BASIS
Most procedures to obtain a highly purified PLA2 involve several purification steps and even different matrixes, which make these procedures more expensive. Extraction is a critical step in detecting desired enzymatic activity in any living organism, especially in those where there is specific interactions with membranes. Designing efficient and selective extraction systems to ensure satisfactory recovery of such desirable activity, and prevent interferences with other enzymes in the assay is a major concern [11, 12]. This results from insufficiently available knowledge about phospholipases existence in marine invertebrates. Thus far, t here has been described a large variety of purification procedures for the PLA2; nevertheless, in the case of complex sources when natural expression of these enzymes is the main source of recovery [13-15], the final yield of many of the proposed procedures is low as a result of multiple steps. Hence, more high-powered procedures are required for the purification of these enzymes, such as affinity chromatography.
To improve interaction during affinity chromatography the ligand nature is, precisely, an important feature, in this case a phospholipid (PC). Phospholipases show significant affinity for this substrate [16, 17]. In the specific case of Condylactis gigantea , soybean PC and egg yolk PC are the best substrates hydrolyzed by this kind of enzyme [18] and the egg yolk PC oxidized is well adapted for affinity-construction support after oxidation procedure. Affinity-based methods for PLA2 taking into account phospholipid ligand immobilization or a similar molecule require calcium for the interaction during fixation [19, 20], and elution of fixed proteins is carried out incorporating EDTA , as used herein.
MATERIALS AND METHODS
Extracts Preparation
The sea anemone Condylactis gigantea was collected at the coast of Havana by specialists from the Faculty of Biology, in Universidad de La Habana. The sea anemone extracts were obtained according to Romero [23].
Chromatographic Partitioning
The initial step for chromatographic partitioning was the same used for isolation and purification of marine toxins from this coelenterate [24].
]]> Chromatographic Procedure of Crude Extracts in Sephadex G-50 GelChromatographic partitioning in thin Sephadex G-50 (Pharmacia Fine Chemicals) was the initial step. About 44-46 mg of proteins, determined by Bradford's method [25], were applied to a column of 3 x 90 cm (preparative scale). Equilibration buffer system was Tris-HCl 50 mmol/L, pH 7,5, CaCl2 40 mmol/L. 3 ml fractions were collected at a flow rate of 30 cm x h-1 (3 mL x min-1).
Affinity Chromatography
An affinity support with oxidized phosphatidylcholine (PC-MANA-Sepharose CL-4B), previously synthetized in our laboratory [26], was used for next chromatographic partitioning. The matrix was equilibrated with the same buffer solution used in the previous purification step and all samples were dialyzed with buffer TrisHCl 0.05 mol/L, pH 7,5, CaCl2 0,04 mol/L (fixation buffer) before application.
Chromatography PC-MANA-Sepharose CL-4B was carried out at 25 °C in a glass column (0.7 x 8 cm). Samples from excluded volume coming from molecular exclusion chromatography (called Peak I G-50) were applied, and the column was exhaustively washed with fixation buffer solution.
As soon as the sample was fixed and the column washed down, elution of fixed proteins was carried out with a buffer solution TrisHCl 0,05 mol/L pH 7.5 containing EDTA 0.04 mol/L. For plotting the elution profile, 3 ml fractions were collected at a flow rate of 90 cm/h in a 1,6 x 5 cm column for the standard procedure, monitored at 280 nm in an Ultrospec 4000 spectrophotometer (Pharmacia Biotech. Sweden).
Due to the highly hydrophobic nature of the immobilized ligand in the gel, a proteins elution profile study of fixed proteins was accomplished in order to check fixed proteins behavior at different flow rates, using a peristaltic pump Pharmacia LKB - Multidrive XL.
Protein Concentration
Protein concentration was determined by Bradford's method [25] using a standard curve with BSA2 mg/mL. For chromatography profiles, proteins concentration was reported from absorbance at 280 nm, which is increased in this type of enzymes [27]. An arbitrary extinction coefficient ξ = 1mL/mg [28-30] was assumed. For data processing and comparisons depending on proteins concentration, values are taken with the same procedure.
Qualitative Assay of Phospholipase Activity
]]> Taking into account the conditions of this assay, Phospholipase A2 activity was corroborated according to León et al [31], which was modified by Bárcenas [18], using both egg yolk PC purified [32] and the fluorogenic substrate 1-palmitoil2-NBD-C12-PC (Avanti Polar Lipids Inc., Alabaster, AL, USA).The presence of the fatty acid or the labeled fatty acid in the TLC confirmed this enzymatic activity [33]; both were compared to standards obtained by hydrolysis of the fluorogenic substrate with the Crotalus durrisus terrificus PLA2 (Sigma-Aldrich, St. Louis, MO, USA).
The Thin Layer Chromatography (TLC)-based method was carried out on a Silica Gel 60 F-254 20x20 plate (Merck, Darmstadt, Germany), using two solvents mixtures:
First solvents mixture: chloroform: methanol: H2O (65: 35: 2.5) until 10 cm high.
Second solvents mixture: oil ether: ethyl ether: glacier acetic acid (70: 30: 1).
All solvents were analytically graded (Sigma Chemicals Co).
Lipids are revealed instantly in an iodine camera and, afterward, they are permanently revealed with the phosphomolybdic reagent [34]. In brief, TLC is sprayed with the reagent at 100°C for 1 h.
León et al. [31] phospholipase activity assay modified by Bárcenas [18], and tested with the fluorescent commercial phospholipid allows visualization of migration without revealing. The fluorescent substrate and product were observed under a UV lamp. As for control, the above-mentioned treatments were performed without Condylactis gigantea mixture.
Polyacrylamide Gel Electrophoresis
Polyacrylamide gel electrophoresis [35] was carried out in a vertical chamber with a 15 % stacking gel and 12,5 % resolving gel. 20 µL of samples previously concentrated and desalinated were applied, using blue bromephenol as 2-mercaptoethanol-free indicator. Molecular weight standards were BSA (66 000 Da), Ovalbumin (45000 Da), Pepsin (34 700 Da), Trypsinogen (24 000 Da), ß -lactoalbumin (18 400 Da), Lysozyme (14 300 Da) and Aprotinin (6 500 Da) purchased from Sigma Chem. Co.
]]> Electrophoretic running parameters were: 100 V, 50 mA y 12 watt for 30 minutes during samples concentration in concentrator gel, and a current of 150 V, 100 mA and 12 watt for 1:07 h was applied to separator gel by a power supply Pharmacia LKB- Multidrive XL.Tincture was completed using Coomasie blue R-250 at 0,25 % in methanol 25 %, acetic acid 10 % and water (V/V/V) during 20 minutes. Colorant excess was eliminated by means of washes during 24 hours with the methanol-acetic acid- water with the previously described proportions, and gel was kept into this solution for preserving.
Statistical Analysis
XLSTAT and STATGRAPHICS Centurion XV were used to determine media and standard deviation in all measurements (at least 3 replicas for each tabulated point) as well as the tests performed by media comparison between both matrixes.
RESULTS AND DISCUSSION
Evaluation of the Purification Process
The purification process was controlled by proteins concentration in each step due to difficulty to continuously detect specific phospholipase activity A2 (analyzed in the bibliographical review) . Consequently, yields obtained are not completely satisfactory and they might be higher if a specific method is used in natural cofactors.
Values of absorbance at 280 nm are related to a significant number of aromatic residues, with the highest values of absorbance at 280 nm peak A1 from affinity, which could indicate the amino acidic composition nature of the proteins fixed to the support, a feature of PLA2 [27], but not specific.
Protein concentration for affinity profiles was followed by absorbance at 280 nm choosing an arbitrary extinction coefficient of 1 mg/mL due to low protein quantities calculated by Bradford. These proteins are characterized by the presence of a high quantity of aromatic residues for molecule, ranging between 13-18 [27]. This fact greatly influences the typical absorbance reading at 280 nm for these amino acids, allowing a sensitive detection tool in front of low protein quantities.
]]> The application of the snake poison from Crotalus durisus terrificus (a known source of PLA2) [21], offered a typical profile for this chromatography and confirmed a similar one with Condylactis Gigantea.Due to complexity to determine and/or quantify the specific enzymatic activity by continuous method for those enzymes, the proteins yield in each step was one of the useful parameters frequently followed. In this case, it was measured in both, continuous (table 1) and batch system (table 2). For statistical analysis, protein concentration was determined by Bradford [25], after 9 chromatographic runs with 3 replicas for each one.
TABLE 1. TOTAL PROTEINS RECOVERED AFTER EACH PURIFICATION STEP IN A CONTINUOUS SYSTEM (DIRECT SYSTEM)
|
| Total proteins (mg) | Yield (%) |
| CE (crude extract) | 148,05 +/- 6,78 | 100 |
| ]]> Eluted fraction 1 (Sephadex G-50) | 77,22 +/- 8,79 | 52,16 +/- 5,92 |
| Affinity chromatography | 28,83 +/- 4,77 | 19,47 +/- 3,22 |
TABLE 2. TOTAL PROTEINS RECOVERED AFTER EACH PURIFICATION STEP BY INDIRECT METHOD (BATCH SYSTEM)
|
| Total proteins (mg) | ]]> Yield (%) |
| CE (crude extract) | 0,095 ± 0,006 | 100 |
| Eluted fraction 1 (Sephadex G-50) | 0,047 ± 0,003 | 49,5 ± 3,2 |
| Affinity chromatography | 0,031 ± 0,002 | 32,4 ± 3,0 |
During continuous system, the sample contacts the support only once, interacting with the same bead during a time limited by flow rate, favoring in this way the strongest interactions among those of affinity. This includes intrinsically those of hydrophobic interactions given the nature of the active center of these enzymes as well as for its raised aromatic amino acids composition. On the other hand, the calcium, in the way to the ideal concentrations for the enzymatic activity of this enzyme determined by Bárcenas [18], favors the activation and shape of the enzyme to fits substrate. Then, exhaustive washes promote elimination of the weak interactions that can survive after this process of interaction of the sample applied to the support. Thus, batch system may be efficiently used when no highly purified PLA2 is needed while continuous system is recommended for highly purified PLA2.
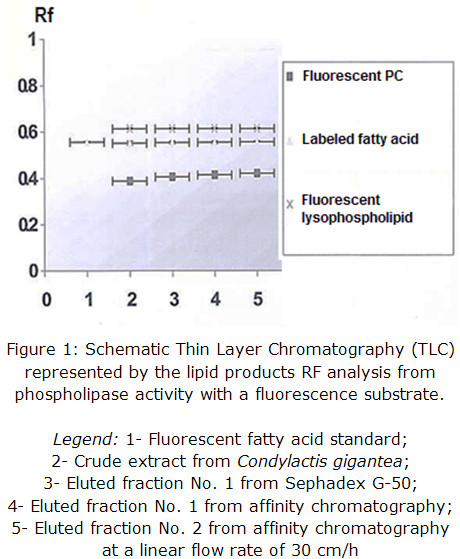
Enzymatic activity from the pools of collected fractions was determined qualitatively in each step of the purification procedure and corroborated for all eluted fractions, being the most intense the one coming from affinity support. This result validates that the obtained protein presents PLA2 activity and it is shown in figure 1 through Rf representation.
The presence of fluorescent lysophospho lipids may be slightly produced by a concomitant phospholipase A1 activity of, at least, one protein component and / or spontaneous hydrolysis due to samples preparation procedure, which could be further determined by fluorescence quantification, if compared to a pattern.
Different profiles were obtained changing flow rates, as shown in figure 2.
4 different flow speeds were checked in order to verify fixed proteins interactions to the chromatographic support. Elution pattern is shown in figure 3. As observed, when reducing flow speed, there is a better resolution of the fractions, which could be explained for differential hydrophobic interactions of proteins with the immobilized ligand in the chromatographic support. This could be confirmed with further assays.
An SDS-PAGE was also a purity criterion in the different steps of the purification process, and it was also taken into account to determine the PM of the proteins fixed to the affinity support.
SDS-PAGE results are shown in figure 3.
]]> A major intense band with a similar mobility to the standard of ß-lactoalbumin (18400) is observed as well as two bands of minor intensity between this standard and lysozyme one (14300).After 3 chromatographic runs with Condylactis gigantea, the total volume of collected fractions from eluted peaks were concentrated with PEG 6000. Proteins electrophoresis in polyacrylamide gels showed three bands with molecular weights at the range between 1400020000 Da; there was a clearly visible one and the other two bands were still very tenuous. These match with those informed for the secreted PL A2 from mammals, snakes and vertebrates in general [36-39].
This element may support the hypothesis that more than one purified proteinaceous components by this procedure from Condylactis gigantea extract shows PLA2 activity, possibly isoforms. This situation is not slightly frequent, since the presence of PLA2 isoforms is typical in these poisons [21, 40]. Similar results have been recently proved by Romero et al, [39] after ion exchange chromatography on CM-Sepharose, a second chromatography step of hydrophobic interaction column on Phenyl-Sepharose and a third purification step in a C18 reverse phase column in HPLC. They founded a highly puri?ed enzyme named CgPLA2 (with a major peak (99.8%) when analyzed by mass spectrometry). This enzyme had a molecular weight of 14.500 for the monomer in the presence of reducing agents, and a little amount of possible isoforms of this enzyme, with molecular masses of 14 269 Da, 14 277 Da and 14 306 Da showed to be identical to the main enzymatic component, CgPLA2, after sequencing of the 32 first N-terminal aminoacid residues [39].
After chromatographic runs at different flow rates, it was observed the diminishing of the main absorption maxima obtained as flow rate raise up, correlated with the appearance of others up to a total number of 3. This is possible due to the effect of a differential affinity of isoforms towards the ligand immobilized in the support. In order to confirm this possibility, it would be necessary to separate the different fractions, and determine the Km for each one of these components presenting this specific activity and/or amino acid sequencing.
A two-step purification procedure for PLA2 is rarely reported. For example, Zhang et al. [41], reported that Lp-PLA2 could be purified rapidly and conveniently through a one-step or a two-step procedure but expressing and purifying recombinant Lp-PLA2 in different heterologous expression systems (P. pastoris and baculovirus, respectively).
This easy and not expensive purification procedure of PLA2 allows the purification of at least one PLA2 from Condylactis gigantea with an acceptable purity for such a variety of biotechnological and pharmacological uses. This affinity procedure avoids presence of many other peaks such as those obtained by DEAE Sephadex A-50 [42].
CONCLUSIONS
The two-step procedure allows purification of three protein components with molecular weights between 18000 and 14000 Da, with at least one of the components possessing phospholipase A2 activity.
]]>ABBREVIATIONS USED
PLA2, phospholipase A2; sn, stereospecific numbering; PC, phosphatidylcholine; sPC, soybean PC; eyPC, egg yolk PC; MANA, monoaminoethyl-N-aminoethyl; EDTA, ethylenediaminetetraacetic acid; UV, ultraviolet; PAGE, poly-acrylamide gel electrophoresis; SDS, sodium dodecyl sulphate; TLC, Thin Layer Chromatography; NBD, 7-nitro-2-1-3-benzoxadiazole-4-amino.
REFERENCES
1. VOLWERK, J. J., LOST, P. C., DE HASS, G. H., GRIFFITH, O. H. "Activation of Porcine Pancreatic Phospholipase a by the Presence of Negative Charges at Lipid-Water Interfase", Biochemistry , 1986, 25, 1726-1733.
2. BERNER, D. L., HAMMOND, E. G. "Phylogeny of Lipase Specificity", Lipids, 1970, 5(6), 558-562.
3. SIX, D. A., DENNIS, E. A. "The Expanding Superfamily of Phospholipase A2 Enzymes: Classification and Characterization", Biochim. Biophys. Acta, 2000, 1488, 1-19
4. MINCHIOTTI, M., SCALAMBRO, M. B., VARGAS, L., CORONEL, C., MADOERY, R. "Isolation of Phospholipase A2 from Soybean (Glycine Max) Seeds: The Study of Its Enzymatic Properties", Enzyme and Microbial Technology, 2008, 42(5), 389-394.
5. MADOERY, R., MINCHIOTTI, M. "Cibacron Blue-Eupergit, an Affinity Matrix for Soybean (Glycine Max) Phospholipase A2 Purification", Enzyme and Microbial Technology, 2006, 38(7), 869-872.
6. CAMPBELL, L. D. "Function-Location-Control", Journal of Science, 1990, 250, 1541.
7. NEVALAINEN, T. J., PEURAVUORI, H. J., QUINN, R. J., LLEWELLYN, L. E., BENZIE, J. A. H., FENNER, P. J., WINKEL, K. D. "Phospholipase A2 in Cnidaria", Comp. Biochem. Physiol., 2004, 139 Part B, 731-735.
8. KNOTZ, S., BOERSMA, M., SABOROWSKI, R. "Microassays for a Set of Enzymes in Individual Small Marine Copepods", Comp Biochem Physiol A Mol Integr Physiol, 2006, 145(3), 406-411.
]]>9. PARK, J., CHO, S. Y., CHOI, S. J. "Purification and Characterization of Hepatic Lipase from Todarodes Pacificus", BMB reports, 2008, 41(3), 254-258.
10. PERERA, E., MOYANO, F. J., DÍAZ, M., PERDOMO-MORALES, R., MONTERO-ALEJO, V., ALONSO, E., CARRILLO, O., GALICH, G. S. "Polymorphism and Partial Characterization of Digestive Enzymes in the Spiny Lobster Panulirus Argus", Comp Biochem Physiol B, 2008, 150(3), 247-254.
11. MALA, J. G., EDWINOLIVER, N. G., KAMINI, N. R., PUVANAKRISHNAN, R. "Mixed Substrate Solid State Fermentation for Production and Extracing Lipase from Aspergillus Niger Mtcc 2594", J Gen Appl Microbiol, 2007, 53(4), 247-253.
12. TALUKDER, M. R., SUSANTO, D., FENG, G., WU, J., CHOI, W. J., CHOW, Y. "Improvement in Extraction and Catalytic Activity of Mucor Javanicus Lipase by Modification of AOT Reverse Micelle", Biotechnol J., 2007, 2(11), 1369-1374.
13. IIJIMA, N., NAKAMURA, M., VENATSU, K., KAYAMA, M. "Partial Purification and Caracterization of Pla2 from Hepatopancreas of Red Sea Bream", Nippon Suisan Gakkaishi, 1990, 56(8), 1331-1339.
]]>14. VERGER, R., FERRATO, F., MANSBACH, C. M., PIERONI, G. "Novel Intestinal Phospholipase A2: Purification and Some Molecular Characteristics", Biochemistry, 1982, 21(26), 6883-6889.
15. ZHANG, Y., XU, T., CHEN, Q., WANG, B., LIU, J. "Expression, Purification, and Refolding of Active Human and Mouse Secreted Group Iie Phospholipase A2", Protein Expression and Purification, 2011, 80(1), 68-73.
16. LESLIE, C. C. "Properties and Regulation of Cytosolic Phospholipase A2", The Journal of Biological Chemistry, 1997, 272(27), 16709-16712.
17. BUNT, G., VAN ROSSUM, G. S., VAN DEN BOSCH, H., VERKLEIJ, A. J., BOONSTRA, J. "Regulation of Cytosolic Phospholipase A2 Activity", Biochemistry, 2000, 39, 7847.
18. BÁRCENAS, J. "Estudio De Algunas De Las Características Funcionales De Una Fosfolipasa a De La Anémona Marina Condylactis Gigantea", Research University of Havana, 1990.
]]>19. VERGER, R., MIERAS, M. C. E., DE HAAS, G. H. "Action of Phospholipase A at Interfaces", J. Biol. Chem., 1973, 248, 4023-4034.
20. DEL MONTE-MARTÍNEZ, A., GONZÁLEZ-BACERIO, J., ROMERO, L., ARAGÓN, C., MARTÍNEZ, D., CHÁVEZ, M. D. L. Á., ÁLVAREZ, C., LANIO, M. E., GUISÁN, J. M., DÍAZ, J. "Improved Purification and Enzymatic Properties of a Mixture of Sticholysin I and II: Isotoxins with Hemolytic and Phospholipase A2 Activities from the Sea Anemone Stichodactyla Helianthus", Protein Expression and Purification, 2014, (0).
21. ROCK, C. O., SNIDER, F. "Rapid Purification of Phospholipase a from Crotalus Adamantheus Venom by Affinity Chromatography", Journal of Biology, 1975, 250, 6564-6566.
22. BURKE, J. E., DENNIS, E. A. "Phospholipase A2 Biochemistry", Cardiovasc. Drugs Ther., 2009, 23, 49-59.
23. ROMERO DEL SOL, D. L., LUBERTA, A., DÁVILA, L., BARRAL, A. M., GARATEIX, A., MAS, R., CHÁVEZ PLANAS, M. D. L. Á. "Aislamiento Y Puri ? cación Parcial De Cuatro Polipéptidos a Partir De La Anémona Marina Condylactis Gigantea", Rev. Biol., 1987, 1, 3-13.
24. ROMERO DEL SOL, D. L. Aislamiento Y Purificación De Toxinas a Partir De La Anemona Condylactis Gigantea. University of Havana, 1997.
25. BRADFORD, M. M. A "Rapid and Sensitive Method for the Quantification of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding", Anal. Biochem., 1976, 86, 248-254.
26. GUISADO-BOURZAC, F., D.L., ROMERO DEL SOL, J.M., GUISÁN SEIJAS, J., GONZÁLEZ-BACEIRO, J., DÍAZ BRITO, A., DEL MONTE-MARTÍNEZ. (2016). "Obtaining an Affinity Support for Phospholipase A2 Isolation and Purification", Revista Cubana de Química, 28(2), 595-609.
27. ALAGÓN, A. C., MOLINER, R. R., POSSANI, L. D., FLETCHER, P. L., CRONAN, J. E., JULIA, J. I. "Purification and Characterization of the Phospholipase A", Biochem. J., 1980, 185, 695-704.
28. ABELSON, J. N., SIMON, M. I. Methods in Enzimology. Edtion ed.: Academic Press, 1990.
29. SCOPE, R. Protein Purification. Principles and Practice.. Edtion ed. New York SpringerVerlag, 1984.
30. LEHNINGER, A. Principles of Biochemistry. Edition ed., 2006.
31. LEÓN, O. S., HENRÍQUEZ, D. R., DÍAZ BRITO, J. "Algunas Características Químico Fisicas Del Tritón X-100 Y Su Relación Con La Fosfolipasa a De La Gorgonia Plexaura Homomalla", Boletín de Inf. Científica del I.Q.B.E. (ACC) , 1984, 3 (2), 24-34.
32. SINGLETON, W. S., GRAY, M. L., WHITE, J. L. "Chromatografically Homogeneus Lecithin from Egg Phospholipids", The Journal of American Oil Chemistry, 1965, 42, 53-56.
33. ABE, A., KELLY, R., SHAYMAN, J. A. "The Measurement of Lysosomal Phospholipase A2 Activity in Plasma", J. Lipid Res., 2010, 51, 2464-2470.
34. DÍAZ, J., ESPINOSA, G., GARCÍA, M. E., ALONSO BIOSCA, M. E., GÓMEZ, T., ROMERO DEL SOL, D. L. Bioquímica Experimental. Editorial Pueblo y Educación, 1989.
35. LAEMMLI, U. K. "Cleavage of Structural Proteins During the Assembly of the Head of Bacteriophage T4", Nature, 1970, 227, 680-685.
36. VALENTIN, E., LAMBEAU, G. "What Can Venom Phospholipases A2 Tell Us About the Functional Diversity of Mammalian Secreted Phospholipases A2?", Biochimie, 2000, 82(9), 815- 831.
37. WURL, M., KUNZE, H. "Purification and Properties of Phospholipase A2 from Human Seminal Plasma", Biochimica et Biophysica Acta (BBA)-Lipids and Lipid Metabolism, 1985, 834(3), 411-418.
38. SUN, M.-Z., LIU, S., YANG, F., GREENAWAY, F. T., XU, Y. "A Novel Phospholipase A2 from Agkistrodon Blomhoffii Ussurensis Venom: Purification, Proteomic, Functional and Structural Characterizations", Biochimie, 2009, 91(4), 558-567.
39. ROMERO, L., MARCUSSI, S., MARCHI-SALVADOR, D. P., SILVA, F. P., FULY, A. L., STÁBELI, R. G., L, S., GONZÁLEZ, J., SOARES, A. M. "Enzymatic and Structural Characterization of a Basic Phospholipase A2 from the Sea Anemone Condylactis Gigantea", Biochimie, 2010, 92, 1063-1071.
40. GROTENDORST, G. R., HESSINGER, D. A. "Purification and Partial Characterization of the Phospholipase A2 and Co-Lytic Factor from Sea Anemone (Aiptasia Pallida) Nematocyst Venom", Toxicon, 1999, 37(12), 1779-1796.
41. ZHANG, F., DONG, L., CAI, M., SHEN, J., WANG, Y. "Heterologous Expression of Lipoprotein-Associated Phospholipase A2 in Different Expression Systems", Protein Expression and Purification, 2006, 48(2), 300-306.
42. WANG, Y., CUI, G., ZHAO, M., YANG, J., WANG, C., GIESE, R. W., PENG, S. "Bioassay-Directed Purification of an Acidic Phospholipase A2 from Agkistrodon Halys Pallas Venom", Toxicon, 2008, 51(7), 1131-1139.
Recibido: 24/01/2016 ]]> Aceptado: 3/08/2016
MSc. Frenkel Guisado-Bourzac, Facultad de Ciencias Naturales, Universidad de Oriente, Santiago de Cuba, Cuba, fguisado@uo.edu.cu
]]>