RESEARCH
A fermentation process for the production of P50 from Serratia marcescens
Proceso de fermentación para la producción de la proteína P50 de Serratia marcescens
Raúl Espinosa1, María del C Abrahantes1, Jorge Valdés1, Victoria Lugo1, Marta Pupo1, Pedro Aguilar1, Maelys Millares1, Inalvis Herrera1, Danae Arias1, Alexis Musacchio2
]]> 1Biotechnology Development Direction2Division of Chemistry and Physics
Center for Genetic Engineering and Biotechnology, CIGB Ave. 31 / 158 and 190, Cubanacán, Playa, PO Box 6162, Havana, Cuba
ABSTRACT
Serratia marcescens is an opportunistic gram-negative enteric bacterium isolated from the respiratory and urinary tracts in humans. Among the secreted S. marcescens extracellular proteins, the P50 protein is produced in large amounts and plays an important role in the pathogenesis of this organism. To produce this protein, a fermentation process was studied. First, in a 23 factorial experimental design different culture supplement as tryptone, yeast extract and peptone, were studied. In 4 L bioreactors the influence of aeration rate and agitation speed over the P50 production were studied in a 32 experimental design. Finally, the optimal growth conditions were established (28 °C, 400 rpm and 0.5 vvm), at this scale, 9.0 ODu/mL, 62% level of P50 protein expression and 220 mg/L as the higher P50 volumetric production, were obtained.
Keywords: cytotoxic protein, P50, Serratia marcescens.
Serratia marcescens es una enterobacteria gram-negativa oportunista aislada de las vías respiratorias y el tracto urinario en humanos. Entre las proteínas extracelulares segregadas de S. marcescens, esta la proteína P50 la que se produce en mayores cantidades y desempeña un papel importante en la patogénesis de este microorganismo. Para producir esta proteína se ha estudiado un proceso de fermentación. En primer lugar, en un diseño experimental, factorial de 23, se estudiaron cultivos suplementados con triptona, extracto de levadura y peptona. En bioreactores de 4L se estudio la influencia de la aireación y la velocidad de agitación sobre la producción de la P50 en un diseño experimental 32. Por último, se establecieron las condiciones óptimas de crecimiento (28 °C, 400 rpm y 0.5 vvm), a esta escala, se obtuvo el 9.0 ODu/mL, 62% de nivel de expresión de la proteína P50 y 220 mg/L como la mayor producción volumétrica de P50.
Palabras clave: proteína citotóxica, proteína P50, Serratia marcescens.
INTRODUCTION
Serratia marcescens is an opportunistic pathogen in immunocompromised hosts (1) and it is a source of nosocomial infections (2). This Gram-negative enteric bacterium is often isolated from respiratory and urinary tracts in humans. S. marcescens causes a wide spectrum of infections such as pneumonia, meningitis, septicemia, urinary tract infection, endocarditis, conjunctivitis, and wound infection (2-4), but the virulence mechanisms of this organism are poorly understood.
S. marcescens secretes many known extracellular proteins, including chitinase, lecithinase, hemolysin, siderophore, lipase, protease, and nuclease (4, 5). S. marcescens serralysin, is especially produced in the largest amounts from pathogenic clinical isolates being considered to play an important role in pathogenesis of this organism (6-8).
Purified serralysin has been used in in vitro and in vivo animal models (9), and its cytotoxic activity has been also evaluated in tumor cells (10-17). The P50 serralysin from S. marcescens has been also identified with similar above mentioned properties, as previously described (10). To perform further preclinical studies, in this work we studied and established the fermentation process of a particular serralysin expressed in S. marcescens.
]]> MATERIALS AND METHODSMicroorganism
The isolated Serratia marcescens strain CMIB4202 (10) was used to prepare the master and working seed culture banks. The stocks were prepared as described (18), growing the cells at 30 ºC in tryptone soy broth and glycerol 15% (v/v).
Antibody
Polyclonal antibodies generated in sheep against the P50, were used for immunochemical analysis (western blot).
Growth conditions
Factorial design to establish P50 production conditions in bioreactors
A 23 factorial experimental design (19) was used to point out the relationships existing between the P50 protein production (Y1), as the variable response, and the culture supplement as experimental factors. The levels of the three independent factors, tryptone (x1), yeast extract (x2) and peptone (x3) concentrations, which were required to obtain the optimum protein production were settled up as present (1) or not present as (-1). The factorial analysis and the statistical test of variances (ANOVA) were performed with the Statgraphics software, v 5.1. All experiments were conducted in triplicate and carried out in bioreactors.
Influence of aeration rate and agitation speed to the growth conditions and P50 production
The influence of aeration rate in the production of P50 was studied in 4 L bioreactors (Biolafitte, France).
]]> Three aeration conditions (0.5, 1.0, 1.5 vvm) in combination with three different agitation speeds (150 rpm, 275 rpm, 400 rpm), were selected for this study. A 32 factorial experimental design was used for the design of experimental and the resultant data analysis.The data analysis was performed according to the 32 experimental design (two independent factors: aeration rate and agitation speed, and three levels 0.5, 1.0 and 1.5 vvm), taking into account the P50 production and using the selected culture media from the 23 experimental design performed in bioreactors.
Conditions for P50 protein production at 4 L scale
The Serratia marcescens strain CMIB4202 was grown in 400 mL of tryptone soy broth medium from Oxoid (Basingstoke, Hampshire, England) in shake flasks (New Brunswick Scientific Co., USA), at 28 ºC, 250 rpm for 5 h, which were used to inoculate the bioreactors. The culture media in bioreactors was composed by the minimal growth medium (MM) (20) supplemented with 10 g tryptone/L, 10 g yeast extract/ L and 10 g glycerol/L. The fermentation process was carried out in 4 L bioreactors at 28 ºC, 400 rpm, and 0.5 vvm aeration rates.
Analysis
Cell concentrations were determined turbidometrically at 620nm (Erma Optical Works, Japan). The expression level of proteins was determined by densitometry using the denaturing 12.5% SDS-PAGE (21), and the resulting protein band patterns were quantified by densitometry (Molecular Analyst Package, BIORAD, USA). The protein concentration was determined by the modified method of Bradford (22). The immunodetection of P50 was performed by Western blot (23) using polyclonal antibodies generated in sheep, again hast the previously purified recombinant protein (Abrantes-Pérez MC, unpublished results).
RESULTS AND DISCUSSION
Influence of the culture supplement in the P50 production
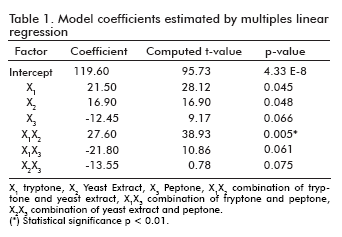
The influence of tryptone, yeast extract and peptone as culture supplements for the P50 production, were studied. The optimal values of selected variables obtained by the regression equations and the calculated coefficients of the model are shown in table 1.
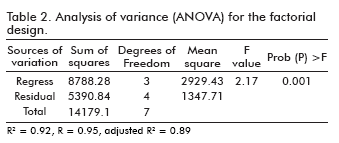
]]> The significance of each coefficient was determined by students t-test and p values, which are listed in table 1. The larger the magnitude of the t-value and the smaller the p-value, the more significant the corresponding coefficient (24). From the calculated values the combination of tryptone and yeast extract gave a direct relationship in the production of P50 protein.The results of the analysis of variables (ANOVA) for the factorial design are given in table 2. The fisher F-test demonstrates a high significance for the regression model (24). The goodness of fit of the model was checked by the determination coefficient (R2). In this case, the value of the determination coefficient (R2= 0.92) indicated that only 8% of the total variation are not explained by the model. The value of the adjusted determination coefficient (adj.R2= 0.89) is also high, which indicates a high significance of the model (24, 25). A higher value of the correlation coefficient (R= 0.95) signifies an excellent correlation between the independent variables.
The application of the methodology (26, 27) yielded the following regression equation which is an empirical relationship between the test variables in coded units:
Y1 = 119.6 + 21.5 X1 + 16.9 X2 -12.45 X3 + 7.6 X1X2 21.8 X1X3 13.55 X2X3
where:
Y1: is the response, which characterizes the P50 production.
X1: represents tryptone.
X2: represents the yeast extract.
X3: represents the peptone. ]]>
X1 X2: represents the combination of tryptone and yeast extract.
X1 X3: represents the combination of tryptone and peptone.
X2 X3: represents the combination of yeast extract and peptone.
The best values in cell growth and production of P50 protein were obtained for culture media contai ning tryptone and yeast extract. The values of expression levels for this protein were similar in this medium, when compared to that contained in its composition peptone and glycerol.
Influence of agitation speed and aeration rate in the P50 production
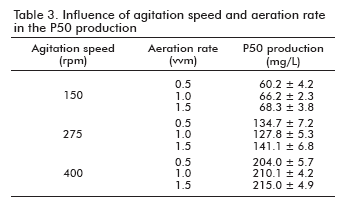
To study the influence of the aeration rate in the production of P50, a set of experiments were carried out in 4 L bioreactors, setting up the agitation speed and varying the aeration rate among 0.5 and 1.5 vvm. The results are shown in table 3. As it can be observed, no influence of the aeration rate in the P50 production was detected, in spite of an increase of the P50 production as a global value, when the agitation speed was increased.
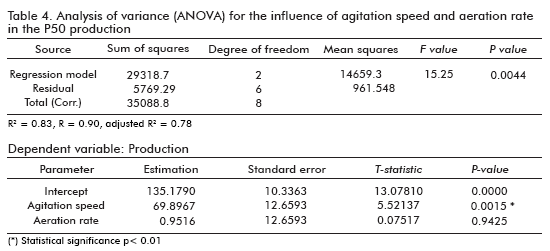
The results of the analysis (ANOVA) (26) of variables over the P50 production gave us a statistical significant influence for the agitation speed (p < 0.0015), as shown in table 4. However, this was not the case of the aeration rate, where no statistical significant difference was observed (p < 0.9425). This analysis of variance for the response variable, P50 production, revealed a very high significance for the regression equation (p model < 0.0044), which confirms the suitability of the linear model. The same results were obtained when the influence of such factors over the cell growth optical densities, were analyzed (data not shown).
Production of P50 protein from Serratia marcescens
]]> The typical schematic fermentation process, for the production of P50 protein at 4 L scale is shown in figure 1. The best conditions for the obtaining of the P50 protein were using 28 ºC, 400 rpm and 0.5 vvm. During the first 10 h of growth an increase in the optical density was observed. The maximum value of the specific growth rate was determined during the first 2 h of growth (μ = 1.2 h-1), being stabilized during the rest of the fermentation process at μ = 0.2 h-1, when an increase in the expression of the P50 protein was detected. The P50 expression levels and the P50 protein production increased during the 10 h fermentation process, reaching the highest production level at this point.At this scale, 9.0 ODu/mL, 62% level of P50 protein expression and 220 mg/L as volumetric production, were obtained. The biomass yield on substrate (Yx/s) and the product yield on substrate (Yp/s) values were 0.22 g biomass/g substrate and 0.002 g product/g substrate, respectively. Previously, it was reported that the maximum optical density obtained for the produc tion of P50 protein from Serratia marcescens was 2.0 ODu/mL (10). In our established conditions a four-fold increased of this parameter was obtained.
The same results were obtained when a comparison in the P50 production values, were performed.
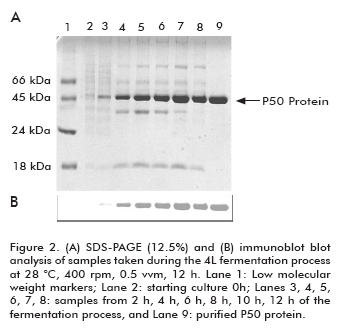
The expression of P50 protein from Serratia marcescens is shown in figure 2. As it can be observed, the P50 protein expression increased during the fermentation process. Immunochemistry analysis of P50 protein from Serratia marcescens obtained during the fermentation process indicated that in spite of the high level of expression, no degradation pattern was observed. As expected, a protein with approximately 50 kDa was obtained and immune-identified by the polyclonal antibodies generated against a previously purified P50 protein.
As a conclusion, a culture media for the P50 production from Serratia marcescens was defined. At the same time, a fermentation process in 4 L laboratory scale was implemented with a good rate of P50 production, having the singularity of no influence of the aeration rate over the P50 protein expression/production. This is the first report where the expression/ production of S. marcescens serralysin was studied.
ACKNOWLEDGEMENTS
We thank Lic. Elizabeth Díaz Lopez for her unconditional help in the preparation of this manuscript.
REFERENCES
]]>1. Goullet P, Picard B. An epidemiological study of Serratia marcescens isolates from nosocomial infections by enzyme electrophoresis. J Med Microbiol 1997;46:1019-28.
2. Hejazi A, Keane CT, Falkiner FR. The use of RAPD-PCR as a typing method for Serratia marcescens. J Med Microbiol 1997;46:913-19.
3. Yu VL. Serratia marcescens: historical perspective and clinical review. N Engl J Med 1979;300:887-93.
4. Aucken HM, Pitt TL. Antibiotic resistance and putative virulence factors of Serratia marcescens with respect to O and K serotypes. J Med Microbiol 1998;47:1105-13.
5. Hines DA, Saurugger PN, Ihler GM, Benedik MJ. Genetic analysis of extracellular proteins of Serratia marcescens. J Bacteriol 1988;170:4141-46.
6. Maeda H. Role of microbial proteases in pathogenesis. Microbiol Immunol 1996;40:685-99.
7. Matsumoto K, Yamamoto T, Kamata R, Maeda H. Pathogenesis of serratial infection: activation of the Hageman factor prekallikrein cascade by serratial protease. J Biochem 1984;96:739-49.
8. Maeda H, Molla A. Pathogenic potentials of bacterial proteases. Clin Chim Acta 1989;185:357-67.
9. Kamata R, Matsumoto K, Okamura R, Yamamoto T, Maeda H. The serratial 56K protease as a major pathogenic factor in serratial keratitis. Clinical and experimental study. Ophthalmology 1985;92:1452-59.
10. Abrahantes-Pérez MC, Reyes-González J, Veliz RG, Bequet-Romero M, Gómez RR, Anais GC, et al. Cytotoxic proteins combined with prodigiosin obtained from Serratia marcescens have both broad and selective cytotoxic activity on tumor cells. J Chemother 2006;18:172-81.
11. Wu J, Akaike T, Hayashida K, Okamoto T, Okuyama A, Maeda H. Enhanced vascular permeability in solid tumor involving peroxynitrite and matrix metalloproteinase. Jpn J Cancer Res 2001;92:439-51.
12. Marty KB, Williams CL, Guynn LJ, Benedik MJ, Blanke SR. Characterization of a cytotoxic factor in culture filtrates of Serratia marcescens. Infect Immun 2002;70:1121-28.
13. Carbonell GV, Amorim CR, Furumura MT, Darini ALC, Fonseca BAL, Yano T. Biological activity of Serratia marcescens cytotoxin. J Med Biol Res 2003;36:351-59.
14. Carbonel GV, Falcon R, Yamada AT, da Fonseca BA, Yano T. Morphological and intracellular alterations induced by Serratia marcescens cytotoxin. Res Microbiol 2004;155:25-30.
15. Chakrabarty AM. Microorganisms and cancer: quest for a therapy. J Bacteriol 2003;185: 683-86.
16. Escobar MM, Carbonell GV, Beriam LOS, Siquiera WJ, Yano T. Cytotoxin production in phytopathogenic and entomopathogenic Serratia marcescens. Rev Latinoam Microbiol 2001;43:165-70.
17. Kida Y, Inoue H, Shimizu T, Kuwano K. Serratia marcescens serralysin induces inflammatory responses through protease-activated receptor 2. Infect Immun 2007;75:164-74.
18. Kirsop BE, Snell JS. Maintenance of Microorganisms. A Manual of Laboratory Methods. Harcourt Brace Jovanovich, Publishers, Academic Press, Inc (London) LTD 1984.
19. Montgomery DC. Design and Analysis of Experiments. John Wiley & Sons (3rd edition), New York 1991.
20. Sambrook J, Fritsch E, Maniatis T. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, New York 1989.
21. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970;227:680-5.
22. Zor T, Selinger Z. Linearization of the Bradford protein assay increases its sensitivity: theoretical and experimental studies. Anal Biochem 1996;236:302-8.
23. Musacchio A, Carmenate T, Delgado M, Gonzalez S. Recombinant Opc meningococcal protein, folded in vitro, elicits bactericidal antibodies after immunization. Vaccine 1997;15:751-8.
24. Akhnazarova S, Kafarov V. Experiment optimization in chemistry and chemical engineering. Mir Publications Moscow 1982.
25. Khuri AI, Cornell JA. Response surfaces: Design and analysis. Marcel Dekker, Inc., New York 1987.
26. Box GEP, Wilson KB. On the experimental attainment of optimum condition. J Roy Stat Soc 1951;B13:1-45.
27. Kenneth WY, Mak Miranda GS, Yap Wah Koon T. Formulation and optimization of two culture media for the production of tumor necrosis factor-b in Escherichia coli. J Chem Tech Biotechnol 1995;62:289-94.
]]> Received in July, 2009.
Accepted for publication in September, 2009.
Raúl Espinosa. Biotechnology Development Direction. Center for Genetic Engineering and Biotechnology, CIGB. Ave. 31 / 158 and 190, Cubanacán, Playa, PO Box 6162, Havana, Cuba. E-mail: raul.espinosa@cigb.edu.cu