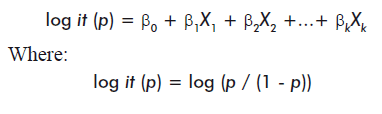
RESEARCH
Estimating the risk for unbalanced chromosomal aberrations in the offspring from translocation-carrying parents
Estimación del riesgo de descendencia con aberraciones cromosómicas desbalanceadas en progenitores portadores de translocaciones
]]>
Joenith Aguilar1, Jorge Bacallao-Guerra2, Jorge Bacallao-Gallestey3, Estela Morales4
1 Departamento de Citogenética, Centro Nacional de Genética Médica. Ave. 31, esq. 146, Cubanacán, Playa, CP 11600, La Habana, Cuba.
2 Departamento de Asistencia Médica, Centro Nacional de Genética Médica. Calle E, No. 309, esq. 15, Plaza de la Revolución, CP 10600 La Habana, Cuba.
3 Departamento de Matemática, Instituto de Cibernética, Matemática y Física, Ministerio de Ciencia, Tecnología y Medio Ambiente, CITMA. Calle Tulipán, esq. Panorama, Plaza de la Revolución, CP 10600, La Habana, Cuba.
4 Centro de Investigaciones y Referencia de Aterosclerosis de La Habana. Ave. 31, esq. 146, Cubanacán, Playa, CP 11600, La Habana, Cuba.
ABSTRACT
Reciprocal and Robertsonian translocations are structural chromosomal aberrations that can produce unbalanced gametes during meiosis. In some cases, these imbalances lead to an offspring with multiple malformations. The purpose of this study was to propose and evaluate a methodology to estimate the risk of live offspring with unbalanced chromosome aberrations (LOUCA) from parents carrying a reciprocal or Robertsonian translocation. The methodology is based on comparing the results from several widely known regression methods: multiple linear, logistic and Poisson regression. Predictive accuracy was evaluated on a database containing information on 41 families of translocations carriers from three Cuban provinces. The results yielded by the three models were quite consistent regarding variable selection (presence of chromosome 9, chromosome 21 and the existence of breaking points in the short arms of the chromosomes involved) and risk estimation. There was a 80% overlap between the classifications produced by the three methods.
]]> Keywords: Risk estimation, translocations, Poisson regression.Las translocaciones recíprocas y robertsonianas son aberraciones cromosómicas estructurales que durante la meiosis pueden originar gametos cromosómicamente desbalanceados. Estos desequilibrios pueden generar una progenie con múltiples malformaciones. El objetivo de este trabajo es proponer una metodología para estimar el riesgo de descendencia con aberraciones cromosómicas desbalanceadas compatibles con la vida, en progenitores portadores de alguna translocación recíproca o robertsoniana. La metodología combina los resultados de los métodos de regresión logística, regresión múltiple y regresión de Poisson. Por sus características, es teóricamente superior a otras variantes descritas en la literatura revisada. El riesgo se estima a partir de una base de datos que contiene información de 41 estudios de familias portadoras de translocaciones, de tres provincias de Cuba. Los resultados con los tres métodos aplicados son coherentes en la selección de las variables predictoras (presencia de cromosomas 9, cromosoma 21 y existencia de puntos de ruptura en los brazos cortos de los cromosomas involucrados) y en las estimaciones del riesgo. El 80% de las familias se clasificaron por estos métodos.
Palabras clave: estimación del riesgo, translocaciones, regresión de Poisson.
INTRODUCTION
The occurrence of reciprocal and Robertsonian translocations during meiosis may produce chromosomal unbalances in the gametes, often leading to offspring with multiple malformations [1, 2]. Therefore, parents from families with known translocations usually need to know the risk or probability of appearance of the consequences of these malformations, which include spontaneous abortion, fetal death and live offspring with unbalanced chromosomal aberrations (LOUCA) [3].
This work proposes a methodology based on the use of generalized linear models for estimating LOUCA risk in the offspring of parents carrying translocations. It has a solid theoretical foundation and can be employed in clinical practice during the process of genetic counseling. Research on this topic dates back to the seventies: see e.g. Daniel in 1979 [4], Stengel-Rutkowski et al. in 1988 [5], Cans et al. in 1993 [6] and Cohen et al. in 1992 [7] and 1994 [8].
Cans et al. proposed, in 1993, the use of logistic regression models for risk estimation, and the use of additive models was also suggested two years later [9].
]]> However, there are theoretical and practical considerations against the use of logistic regression as the sole tool for estimating risk. Like every other linear model, logistic regression is very sensitive to the effects of multi-colinearity when used for explanatory purposes (i.e. the selection of variables relevant to risk estimation). In addition, it assumes the existence of a linear relationship between the dependent variable and the predictors.To further compound matters, data obtained from family studies fails to comply with a basic assumption of logistic regression: that of independence between the subjects. This “family effect” arises because relatives are not only genetically closer than non relatives, but share exactly the same values for the variables describing the translocation inherited throughout the family.
The procedure proposed here sidesteps these obstacles by using a classification into risk groups that is based on the results from multiple and Poisson regressions, thus modifying the structure of data from familial translocation studies. It constitutes a combination of several complementary statistic techniques that can be used to obtain an objective estimation of risk to be later used during genetic counseling.
MATERIALS AND METHODS
Database
The original database contains data from 200 subjects in 41 studies of families carrying reciprocal or Robertsonian translocations. Each family study took into account the individuals with filial relations with respect to the purpose of each study, including this last one. A total of 26 different translocations exhibiting unique breakpoints are present in the data (Figure 1). This database, which constitutes the result of a collaborative effort between the cytogenetic laboratories of Havana, Havana city and Pinar del Río, only included karyotyped, inherited translocations between autosomal chromosomes. Breakpoints were homogenized to a resolution of 400 bands, following the International System for Cytogenetic Nomenclature (ISCN 2005) [10]. The length of centric and translocated segments was measured from the breakpoint to the terminal zone of the long and short arms, with an accuracy of 0.5 mm (G bands). Each family study took into account the individuals with filial relations with respect to the purpose of each study, including this last one.
The offspring status variable, containing six categories (non carrier, balanced carrier, spontaneous abortion, fetal death, neonatal death and LOUCA) was restructured as a binary variable, grouping all conditions excepting LOUCA into a single category. Five chromosomal groups were created, based on the specific chromosome involved in each translocation. Three categories were also created depending on the location of the breakpoint: pp, pq and qq. The parental origin of the translocation was also taken into account. The age of the carrier parent was excluded from the analysis, as recent studies have failed to find a link between age and LOUCA risk [11] and, in addition, a significant portion of the data was missing for this variable. Gamete variability was also excluded, since there were only some cases with LOUCA. Table 1 summarizes the tentative predictors used in the models. The nominal variables chromosomal group and breakpoint location, with five and three categories respectively, were transformed into dummy variables whose values are either 0 or 1 for their inclusion into the regression models.
Statistical analysis
The methodological strategy followed consisted on the application of three regression models, analyzing the agreement between their results. Taking into account the limitations of each procedure and setting aside statistical considerations, this triangulation strategy already improves the reliability of the end result if the same data, analyzed by different methods that part from different sets of assumptions, yield the same outcome. Risk estimation used the same cut-off thresholds employed for genetic counseling (Low risk: < 5; Moderate risk: from 5 to 15%; High risk: > 15%).
The SPSS (Statistical Package for the Social Sciences, version 15.0) software application was used to run the logistic and multiple linear regression models.
]]> The statistical package STATA was used for Poisson regression.Logistic regression
The results were compared by analyzing the data with a logistic regression model (using the 200 subject database). Due care was exercised during the use of this model, as its application for this specific case has some pitfalls.
Logistic regression [12-15] is a specific case of generalized linear models. It is used to model a categorical response from predictors that may be indistinctly continuous or discrete ordinals (in most cases, this response is binary). In this model, p represents the probability of success and the predictors, or independent variables, are represented by Xi:
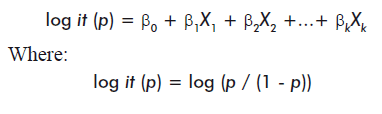
Binary logistic regression is widely used in biomedical and epidemiological research, as the shape of the logistic function is ideal for modeling risk and dichotomic responses, such as the presence or absence of a specific disorder [12].
Multiple lineal regression
This model is described by the equation:
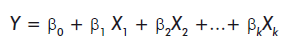
Where Y is the dependent variable and the subscripted X and β represent the independent variables or predictors and the parameters of the model, respectively.
The model was used on a modified version of the original database that was prepared to minimize the “family effect” affecting the logistic regression model described above. In this new database the unit of analysis is the family (families are independent from one another), and the dependent variable (defined in the [0, 1] interval) indicates the proportion of affected individuals within each family. In this manner, individual and family risks are one and the same.
This modification can be introduced without problems because risk, in this dataset, is defined by factors intrinsic to the specific translocation affecting each family, rather than by individual traits. The variables examined in this study that can influence the risk of appearance of LOUCA characterize only the inherited translocation and not other pre-zygotic and post-zygotic genetic phenomena (which can also influence risk) that vary from carrier to carrier. This work, therefore, does not violate the underlying assumptions of its methodology.
Poisson regression
Poisson regression [13-15] is used for modeling counting-type variables and, especially, the risk of appearance of low frequency events. The nature of the problem examined in this work, therefore, lends itself to the use of Poisson regression, as the number of cases in a family is a counting-type variable that can be used for assignation into a risk group.
From a theoretical standpoint, this is the most adequate model (and the results support this reasoning); therefore, its results must be assigned a larger weight when analyzing the results of the classification into risk groups.
]]> A random variable is said to follow a Poisson distribution if its probability function can be written as:
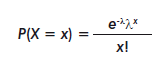
The basic formulation of Poisson regression consists of writing the mean of the counting variable as the exponent of a lineal function of the predictors:
![]()
Due to the resulting structure, this model was applied to the modified database; using the variable Number of LOUCA cases in the family and estimating individual risk as the number of cases in the family divided by family size. Faced with the uncertainty of whether family size might influence the outcome, additional models including this variable were considered and compared to models excluding it. However, family size did not have any significant influence on the results.
RESULTS
Logistic regression
Model 3, predicting LOUCA risk from variables TRAS2 (Gc2), TRAS5 (Gc5) and BRA1 (pp), was obtained after three steps. All three variables are relevant, with signification levels of 0.003, 0 and 0.013, respectively (Table 2). R2 is a coefficient representing to what degree the observed variability is explained by each model. This parameter behaved for each model as follows: Model 1, 0.065; Model 2, 0.166; and Model 3, 0.228. Model 3 is different from the two preceding models. After this study, the subjects were classified according to their LOUCA risk, estimated by logistic regression (See Annex).
]]> Multiple linear regressionA multiple linear regression model was fitted to the data afterwards, using frequency as the dependent variable. The fitting followed the forward method, adding variables as long as R2 increased, and stopping when further inclusions did not result in significant increases of R2.
The best model obtained with the forward method contains the variables TRAS5 (Gc5) and BRA1 (pp) as predictors (adjusted R2 = 0.27, Table 3). These are variables with significant effects (0.004 and 0.002, respectively) (Table 4) that were also selected by the logistic regression model analyzed earlier.
Another variable that might be included in this model would be TRAS2; in this manner, the present model would match the variable set of the logistic model used in the preceding section. After this analysis, the families were classified according to LOUCA risk. No families fell into the moderate risk category when using this model (See Annex).
Poisson regression
Poisson regression was also applied using the family as unit of analysis, and setting the number of LOUCA cases as dependent variable. A new variable denominated family size (TMFA) was included, given that the number of affected cases depends on family size and its addition would prevent, therefore, the occurrence of logical impossibilities such as families with more affected than susceptible individuals. The model was fitted with the robust regression option of the commercially available STATA software package, in order to guarantee that the result is resistant to the effect of outliers. TRAS5 (Gc5), TRAS2 (Gc2) and BRA1 (pp) emerged as relevant variables (Table 5). Since the number of events (variable to be predicted) logically depends on family size, the latter was included as a covariant in the model.
All other things being equal, the variable to predict depended much more on genetic factors than on family size (see the P value of variable TMFA in table 6).
Of note, also, is the fact that using the Poisson regression model resulted, again, in the inclusion of some families in the moderate risk category (see Annex)
DISCUSSION
All three models yielded similar results regarding their most relevant variables and the classification into risk groups, whether using individuals or families as the unit of statistical analysis.
]]> In the three models there are variables indicating the presence of the trait in question that increase risk in a significant manner. One is the presence of breakpoints in the pp arms and another, the involvement of chromosomes 21 and 22. Another variable, which in one case failed to reach statistical significance, was the presence of chromosome 9, included in the models of Poisson and logistic regression. Our results, similar to those of Cans et al. in 1993 [6], highlight the importance of these variables for an understanding of the genetic phenomenon under study.Cans et al. also pointed out that in the case of these chromosomes, risk is even higher when breakpoints are located in both short arms [6]. Our results, despite differences in sample populations and analysis methods, are similar; confirming the validity of our approach and the predictive capacity of the chosen variables. There is a large body of research pointing at chromosome 9 for its frequent involvement in LOUCA, based in the segregation of translocations where it is included [16, 17].
Chromosome 22 is also in Gc5, and therefore is singled out by all models as a risk factor. Although no LOUCA cases involving this chromosome were found in our database, that was not the case for the database used by Cans et al. 1993 [6] due to the high prevalence of t(11;22), which is linked to the presence of this trait in the offspring.
The classification into risk groups was identical among all three models for 80% of the families (see Annex). This result, together with the fact that all models also selected the same variables, further underscores the relevance of the latter as predictors. In general, Poisson regression was more accurate; i.e. it yielded a theoretical risk probability closer to the “observed” values. The model it produced failed only for family 34, where it yielded a risk probability far from the actual value (Annex) since two out of the three members of that family are LOUCA cases. It also identified as significant the presence of Gc2 (chromosome 9), perhaps artificially decreasing the theoretical risk for LOUCA due to the fact that out of the three predictors selected by the Poisson regression (Gc5, Gc2 and “pp”), this is the only one with breakpoints in the short arms of both chromosomes involved in the translocation (“pp”).
Analyzing the predictive accuracy of the Poisson regression, it is possible to see that 1) it was the only model that correctly classified family 40, according to the data. This family had only Gc2 (variable with a value = 1) out of the three predictors selected by the model, demonstrating the significance of the involvement of chromosome 9 in a translocation and its relationship with the probability of LOUCA [18]; and 2) studies of different families with the same translocation yielded different risk probabilities (a phenomenon that had not been observed previously), although they continued to fall within identical risk groups. This is accounted for by differences in the mathematical formulations of the different models, which imply, in the case of the Poisson regression, the use of information unique to each family and, obviously, the introduction of differences between families that share a common translocation.
There is a 95% overlap between the Poisson and logistic regression models regarding the classification of families into risk groups. This overlap decreased to 80% when all three models were considered. The high coincidence between the Poisson and logistic regression models may arise, notwithstanding the characteristics of each one, from the fact that they use the same variables (Gc5, Gc2 and “pp”) as predictors. Regarding multiple regression, it was observed that the presence of breakpoints in the short arms (as in family 29) always produces a high-risk translocation, independently of whether it results in LOUCA or not.
The multiple regression model selected, as predictors, the presence of breakpoints in both short arms (“pp”, BRA1 = 1) and Gc5 (chromosomes 21 and 22, TRAS5 = 1). In addition, none of the translocations in the database has both predictors simultaneously. Given that not many combinations can produce a high-risk classification when using this model, it is sufficient to have one of them for obtaining such a classification, independently from the existence or noof a history of LOUCA in the family. In this model the parameter for variable “pp” is larger than that for variable Gc5; therefore, expected risk is larger when the model is applied to a family with a translocation having both breakpoints in short arms.
Families 29 and 40 were not assigned to the same risk groups by the three models (See Annex). Family 40 should be classified as high risk; however, only the Poisson model identifies it as such, for reasons explained above. Regarding family 29, it should be assigned to the moderate risk group, yet only the logistical regression model does so. It should be noticed that for this last family the value produced by the Poisson regression model came close to that of the moderate risk category. Family 7 was misclassified by all three models as high risk, when it actually does not have a history of LOUCA. This situation was caused by the fact that the translocation of this family contains chromosome 22, which is part of Gc5 (a variable to which all three models assign a large weight) and is absent from all other cases in the database. Therefore, given that chromosome 22 is included in the same group as chromosome 21 (involved in several translocations with a clear link to LOUCA), the family was erroneously assigned to the high risk group.
The three proposed models correctly assign each family to their corresponding risk group, according to the actual data used to verify the accuracy of the prediction. The few observed discrepancies between prediction and reality are considered normal when using empirical estimations that depend on “observed” data [19].
It is important to recap, once again, the main results of the methodology. A technique for estimating the risk for LOUCA was proposed that is based on the use of models. Specifically, these estimations were performed using logistic regression with a database of 200 individual cases, multiple regression in a database of 41 families (corresponding to the previous 200 cases), and Poisson regression in the latter database. Each of these methods has advantages and disadvantages that were discussed above.
]]> Estimating LOUCA risk goes beyond the obtention of a cold number: it entails a careful analysis of the results obtained through all proposed methods and of the conclusions, weighted with the experience of the specialist. The latter adds an important part of rationality to the proposed strategy.ACKNOWLEDGEMENTS
We would like to thank the genetic counselors of the provincial Medical Genetics network from Pinar del Río, and especially, Lic. Olga Luisa Quiñones Masa and Dr. Reinaldo Menéndez García, for their collaboration with this investigation.
REFERENCES
1. Morel F, Douet-Guilbert N, Le Bris MJ, Herry A, Amice V, Amice J, et al. Meiotic segregation of translocations during male gametogenesis. Int J Androl. 2004;27(4): 200-12.
2. Young ID. Risk calculation in genetic counseling. 3rd ed. Oxford: Oxford University Press; 2007.
]]> 3. Stasiewicz-Jarocka B, Haus O, Van Assche E, Kostyk E, Constantinou M, Rybałko A, et al. Genetic counseling in carriers of reciprocal chromosomal translocations involving long arm of chromosome 16. Clin Genet. 2004;66(3):189-207.4. Daniel A. Structural differences in reciprocal translocations. Potential for a model of risk in Rcp. Hum Genet. 1979; 51(2):171-82.
5. Stene J, Stengel-Rutkowski S. Genetic risks of familial reciprocal and Robertsonian translocation carriers. In: Daniel A, editor. The Cytogenetics of Mammalian Autosomal Rearrangements. New York: Alan R Liss; 1988. p. 54-61.
6. Cans C, Cohen O, Lavergne C, Mermet MA, Demongeot J, Jalbert P. Logistic regression model to estimate the risk of unbalanced offspring in reciprocal translocations. Hum Genet. 1993;92(6):598-604.
7. Cohen O, Simonet M, Cans C, Mermet MA, Demongeot J, Amblard F, et al. Human reciprocal translocations: a new computer system for genetic counseling. Ann Genet. 1992;35(4):193-201.
8. Cohen O, Cans C, Mermet MA, Demongeot J, Jalbert P. Viability thresholds for partial trisomies and monosomies. A study of 1,159 viable unbalanced reciprocal translocations. Hum Genet. 1994; 93(2):188-94.
9. Cans C, Lavergne C. De la régression logistique vers un modele additif généralisé: un exemple d´application. Rev Stat Appl. 1995;43(2):77-90.
10. ShafFer LG, Tommerup N. ISCN: An International System for Human Cytogenetic Nomenclature. Memphis: Karger; 2005.
11. McKinlay Gardner RJ, Sutherland GR, editors. Chromosome abnormalities and genetic counseling. 3rd ed. Oxford: Oxford University Press; 2004.
12. Kleinbaum DG, Klein M. Logistic regression. A self learning text. New York: Springer-Verlag; 2002.
13. Agresti A. Categorical Data Analysis. 1st ed. New York: John Wiley & Sons; 1990.
14. McCullagh P, Nelder JA. Generalized Linear Models. 2nd ed. London: Chapman and Hall/CRC; 1989.
15. Christensen R. Loglinear Models. 1st ed. New York: Springer-Verlag; 1990.
16. Sankoff D, Deneault M, Turbis P, Allen C. Chromosomal distributions of breakpoints in cancer, infertility, and evolution. Theor Popul Biol. 2002;61(4):497-501.
17. Harper PS. Practical genetic counselling. 6th ed. Verlag: Hodder Arnold; 2004.
18. Panasiuk B, Danik J, Lurie IW, Stasiewicz-Jarocka B, Lesniewicz R, Sawicka A, et al. Reciprocal chromosome translocations involving short arm of chromosome 9 as a risk factor of unfavorable pregnancy outcomes after meiotic malsegregation 2:2. Adv Med Sci. 2009;54(2):203-10.
19. Nussbaum RL, Mclnnes RR, Willard HF, editors. Thompson and Thompson. Consejo genético y evaluación del riesgo. 7th ed. Barcelona: Elsevier Masson; 2008.
Received in November, 2010.
Accepted for publication in September, 2011.
Jorge Bacallao-Guerra. Departamento de Asistencia Médica, Centro Nacional de Genética Médica. Calle E, No. 309, esq. 15, Plaza de la Revolución, CP 10600 La Habana, Cuba. E-mail:bacallao@icmaf.cu. ]]>