RESEARCH
Design of the purification process by metal-chelate affinity chromatography of a new vaccine antigen for the control of sea lice
Diseño del proceso de purificación mediante cromatografía de afinidad de quelatos metálicos de un nuevo antígeno vacunal para el control del piojo de mar
]]>
Carlos E Perez-Heredia1, Nemecio González-Fernández1, Licette León-Barreras1, Eladio Salazar-Gomez1, Rafael Pimentel-Pérez1, Eulogio Pimentel-Vázquez2, Carmen García-Molina3, Yamila Carpio-González4, Mario P Estrada-García4, Miladys Limonta-Fernández5
1 Grupo de Desarrollo Tecnológico, Centro de Ingeniería Genética y Biotecnología (CIGB). Circunvalación Norte y Ave. Finlay. Camagüey, Cuba.
2 Dirección, CIGB. Camagüey, Cuba.
3 Laboratorio de Análisis, Dirección de Calidad, CIGB. Camagüey, Cuba.
4 Laboratorio de Transgénesis de Peces, Dirección de Investigaciones Agropecuarias, CIGB. Ave. 31 e/ 158 y 190, Cubanacán, Playa, PO Box 6162, CP 10600, La Habana, Cuba. ]]>
5 Departamento de Purificación, Dirección de Desarrollo Tecnológico, CIGB. La Habana, Cuba.
ABSTRACT
The present work aims at designing the purification process of protein MY32/Ls to obtain the active pharmaceutical ingredient (API) against sea lice, marine pathogen that significantly affects the salmon industry in South America and Europe. A compact purification process was designed, based on the analysis of alternatives to establish the steps of rupture, chromatography and renaturation. Due to technical-economic advantages, a chemical rupture process was established using 8 mol/L urea for 1 hour, which was 20 % cheaper than mechanical rupture with glass beads. For washing and elution of the protein in the chromatographic step, pH and imidazole were evaluated. Imidazole was selected for the technical-economic advantages it offered; the costs of the selected variant were 12.33 % lower than washing and elution by pH. In scaling the chromatographic step, an economic variant was assessed with modifications of the urea concentration in washing and elution, lowering the costs of the step 3.4 %, which was moreover attractive for its impact on reducing volumes to be handled in the renaturation step. Tween 80 was selected in the renaturing step, which allowed obtaining the protein in solution. The obtained API induced higher antibody titers in C57Bl/6 mice than the laboratory standard used (three times more on average). The API specifications satisfy the requirements for the formulation of the new vaccine against sea lice.
Keywords: protein purification, sea lice vaccine, MY32/Ls antigen.
RESUMEN
Este trabajo tuvo como objetivo el diseño de un proceso de purificación de la proteína MY32/Ls para obtener el Ingrediente Farmacéutico Activo (IFA) contra el piojo de mar, patógeno marino que afecta considerablemente la industria salmonera de Suramérica y Europa. El proceso se diseñó compacto, basado en el análisis de alternativas para establecer las etapas de ruptura, cromatografía y renaturalización. Por sus ventajas técnico-económicas previstas, se estableció un proceso de ruptura química con urea 8 mol/L durante 1 hora, con un 20 % de ahorro respecto a la ruptura mecánica con perlas de vidrio. Para el lavado y la elución de la proteína se evaluó el pH y el imidazol en la etapa cromatográfica. Se escogió el imidazol por las ventajas técnico-económicas brindadas, con costos de la variante seleccionada inferiores en un 12.33 % a los del lavado y elución por cambios de pH. En el escalado de la etapa cromatográfica se evaluó una variante económica con modificaciones en la concentración de urea en el lavado y elución, con costos reducidos en un 3.4 %, más atractiva por el manejo de volúmenes menores en la etapa de renaturalización. En esta última se seleccionó el Tween 80, lo cual posibilitó obtener la proteína en solución. El IFA obtenido indujo títulos de anticuerpos IgG en ratones C5Bl/6 mayores que los del patrón a escala de laboratorio, tres veces como promedio. Las especificaciones del IFA obtenido satisfacen los requerimientos para la formulación de una nueva vacuna contra el piojo de mar.
Palabras clave: purificación de proteína, vacuna contra el piojo de mar, antígeno MY32/Ls.
]]>
INTRODUCTION
In the last three decades the salmon industry has been greatly affected by sea lice, marine pathogen (ectoparasite) that considerably affects marketing of these fish in the world market. The main effect is caused by the genera Pseudocaligus, Caligus, Lepeophtheirus [1, 2]. Areas dedicated to intensive salmon culture reported over 100 million dollar losses annually due to the presence of lice in their cultures [3].
Carpio et al. [4, 5] obtained two clones of Escherichia coli (BL21 DE3) by recombination, expressing proteins MY32Cr and MY32/Ls as inclusion bodies. The proteins are active on the digestive system of sea lice and therefore limit their development and reproduction, eventually causing death.
The MY32/Ls protein has an approximate size of 22 kDa and an amino acid sequence that retains only some areas or regions similar for both [6]. A tail of six histidines (His6) was fused to its amino acid sequence favoring affinity purification with metal ions. Taking into account sea-lice damages and the opportunity to create a unique product for the European market, a purification process was designed to obtain the API of MY32/Ls protein.
MATERIALS AND METHODS
Obtaining the biomass
]]> To carry out the studies shown in this paper, biomass obtained from a strain of E. coli (BL21 DE3) transformed with a fragment of the akirin gene of Lepeophtheirus salmonis (GenBank accession number: ADD38399) was used. This fragment presented 34-38 % identity to the sequence encoding the akirin protein reported for other arthropods [4, 5]. Complementary DNA encoding protein MY32/Ls was cloned into the pET28 expression vector under the T7 phage promoter, which allows regulating the expression of the gene of interest by adding IPTG to the culture medium.Strategy for purification of the inclusion bodies
Rupture step
Two rupture methods were evaluated: mechanical rupture and chemical rupture. In the first case a ball-mill with glass beads of 0.5 mm diameter was used in the presence of 0.1 mol/L phosphate buffer and 0.1 % Triton X-100 at pH 7. In the case of the chemical rupture only a chaotropic solution composed of 0.1 mol/L phosphate buffer and 8 mol/L urea with pH 8 was used. In the evaluation of both ruptures, the biomass/rupture buffer ratio was set at 1:2 (w/v) (approximately 330 g/L). For mechanical rupture, the ratio of biomass suspension/glass beads was approximately 1:1 (v/v). Protein determination and SDS-PAGE were carried out on the rupture supernatants.
Metal ion affinity chromatography
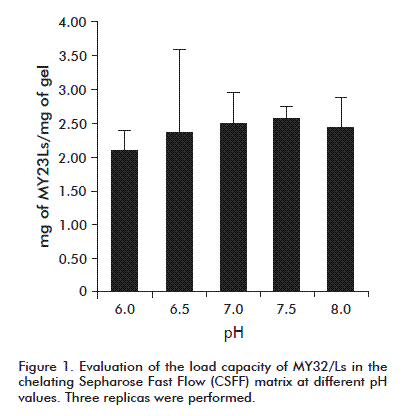
A chelating Sepharose Fast Flow (CSFF) matrix was used; column size was changed depending on the experiment. A study was conducted to evaluate the load capacity of the column at different pH values. It began with a known amount of initial protein and affinity matrix. Phosphate buffer (0.1 mol/L and 8 mol/L of urea were used. pH values evaluated were 6, 6.5, 7, 7.5, 8. The load capacity was determined by the difference in the concentrations of total proteins and SDS-PAGE was performed for subsequent densitometry analysis.
Evaluation of washing and elution by pH
The operation of an XK50 column with 55 mL of CSFF was evaluated in the chromatographic step by performing washing and elution with pH variation. The operational parameters were the following: injection: 10.2 cm/h and washing/elution: 25 cm/h. The pH was lowered using different buffer solutions and the final elution was performed with 0.5 mol imidazole/L to remove remnants of matrix-anchored proteins. The buffers used and the experiment design corresponded to those suggested by the manufacturer of the matrix [7]: equilibrium, 3 column volumes of 0.1 mol/L phosphate buffer (13.58 g/L Na2HPO4, 0.60 g/L NaH2PO4 • H2O) and 8 mol/L of urea at pH 8; washing: 3 column volumes of 0.1 mol/L phosphate buffer (3.58 g/L Na2HPO4, 10.32 g/L NaH2PO4 • H2O) and 8 mol/L of urea at pH 6.3; elution: 5 column volumes of 0.1 mol/L phosphate buffer (13.8 g/L NaH2PO4 • H2O) and 8 mol/L urea at pH 4.5 and elution B: two column volumes of 0.04 mol/L phosphate buffer (3.32 g/L Na2HPO4, 2.29 g/L NaH2PO4 • H2O) and 8 mol/L urea, 0.5 M NaCl and 0.5 M imidazole at pH 7. The total protein concentration was determined, followed by an SDS-PAGE for densitometry analysis.
Evaluation of washing and elution using imidazole
To establish the imidazole concentrations to be used in the chromatographic step, a column of 0.5 mL CSFF was used; washing and elution were performed by ion competition using imidazole. The operational parameters were as follows: injection: 10.2 cm/h and washing/elution: 25 cm/h. Imidazole concentration was increased stepwise fol-lowing the sequence: 0, 0.002, 0.005, 0.01, 0.02, 0.03, 0.04, 0.05, 0.1, 0.3 and 0.5 mol/L. The composition and pH of the buffers used correspond to those reported by the manufacturer of the matrix: 0.1 mol/L phosphate buffer, 8 mol/L urea and imidazole (as corresponds) at pH 7.
]]> Later, using an XK50 column with 55 mL CSFF, the selected imidazole concentrations were evaluated, 3 column volumes were used for washing and for elution, plus the fixed column load value. Sampling was done every 2 column volumes for washing and elution. Total protein concentration was determined and SDS-PAGE was performed for subsequent densitometry analysis.Scaling the chromatography step
Two variants (“baseline” and “economic”) were evaluated, in presence of 8 mol/L urea and another in 4 mol/L. The evaluation was performed maintaining a constant linear speed (injection: 10.2 cm/h; washing/elution: 25 cm/h) with 300 mL of matrix packed in an XK50 column at a temperature of 22 °C. The runs were performed at 22 °C (room temperature). The experiment was followed by measuring absorbance at 280 nm, which would allow better adjustment of work volumes. To carry out the experiment, 3 column volumes were used for washing and elution composed of 0.1 mol/L phosphate buffer and urea: 4 mol/L or 8 mol/L (as appropriate) at pH 7 and the imidazole concentrations established in the previous experiment. Total protein concentration was determined and SDS-PAGE was performed for subsequent densitometry analysis.
Renaturation and dialysis
Evaluation of renaturation using Tween 20 and 80
The presence of Tween 20 and Tween 80, as additives to prevent aggregation of protein MY32/Ls during the renaturation step was evaluated. This assay was performed using the renaturation dilution method. It was ensured that the initial protein concentration was below 1 mg/mL. Tween 20 or Tween 80 ratios for 0, 0.2, 0.4, 1.2 and 2 mg per mg of total proteins were evaluated. The supernatant of each of the variants was analyzed for total protein concentration.
Renaturation using tangential flow filtration
The use of polyethersulfone ultrafiltration cassettes (PESU) of 8 kDa (Slice, with an area of 0.1 m2) was determined for evaluation of this step. Experiments were carried out using a Sartorius SartoJet Pump (with Holder Sartoflow Alpha) ultrafiltration system. As starting material, eluate from the chromatographic step (economic variant) was used. Buffer composition for renaturation/diafiltration was 0.05 mol/L phosphate buffer and 0.01 mol/L EDTA with pH 5.
Renaturation was performed by adding the Tween 80 calculated in the first 0.5 volume of renaturation/diafiltration buffer. Renaturation was concluded by passing another 0.5 volume of the same buffer. Subsequently, the renatured protein solution was concentrated to 2.5 mg/mL total proteins. The renatured and concentrated protein solution was dialyzed against 4 volumes of renaturation/diafiltration buffer. The concentration of total proteins was determined and SDS-PAGE was performed for subsequent densitometric analysis.
Analytical techniques
]]> The bicinchoninic acid (BCA) technique was used to determine total protein concentration. Samples were incubated at 37 °C for 30 min and reading was carried out at 562 nm. The calibration curve was performed using a BSA standard at a concentration of 1 g/L.SDS-PAGE (12.5% acrylamide concentration) was done under reducing conditions using 15 μg of total protein per well [8]. Western Blotting was carried out as described by Joy-Schaffer et al. [9] using the anti-His6 antibody (dilution 1/4000).
Evaluation of MY32/Ls immunogenicity in mice
Immunization
C57BL/6 female, 12-weeks-old, mice were immunized. Animals were divided in three groups according to the immunogen administered in 250 μL by intraperitoneal route on days 0 and 14: I) control group, mice immunized with phosphate buffered saline (PBS); II) mice immunized with 40 μg of an analytical standard of MY32/Ls adjuvanted with Montanide 888 VG; and III) mice immunized with 40 μg of the MY32/Ls active pharmaceutical ingredient (API) formulated in Montanide 888 VG. The MY32/Ls standard was obtained at laboratory research scale according to transfer processes to the technological development phase of the project. Animals were tested on days 0, 14, 28 and 58.
Antibody response determinations
The presence of antibodies in the sera of immunized animals was determined by an indirect ELISA system. High binding ELISA 96 wells plates (Costar, USA) were coated with 100 μL/well of 0.005 g/L MY32/ Ls recombinant protein produced in E. coli and purified (> 90 % purity), dissolved in coating solution composed of 1.6 g/L Na2CO3, 2.9 g/L NaHCO3 at pH 9.5 in PBS, and wells were incubated for 6 h at 4 °C. The plates were incubated in a humid chamber. Subsequently, the plates were washed three times with PBS-0.5 g/L Tween 20 (PBS-T) washing solution and blocked with 200 μL/well of blocking solution (50 g/L skim milk dissolved in PBS-T) for 1 h at 37 °C to block any non-specific binding. Later, the plates were washed three times with PBS-T washing solution and 100 μL/well were added of serial ½ dilutions of sera prepared in blocking solution (initial dilution of 1:200).
The plates were incubated 2 h at 37 °C, further washed four times with PBS-T, and 100 μL/well of anti-mouse IgG peroxidase conjugate (Sigma) prepared in blocking solution at a 1:20 000 dilution were added, followed by incubation for 1 h at 37 °C. After four washings with PBS-T, 0.00042 mol/L 3’,3’,5,5’- tetrametilbenzidine (TMB) and 0.03 % H2O2 in substrate buffer (20 g/L Na2HPO4, 10 g/L citric acid solution, pH 5.0) were added and incubated 5 min at 25 °C.
The reaction was stopped by adding 50 μL/well of a solution of 125 g/L H2SO4 and the absorbance was determined at 450 nm in a plate reader (Sunrise-Basic Tecan, Austria). Serum titers were defined as the reciprocal of the highest dilution for which absorbance values, corresponding to serum dilutions were greater than twice the absorbance obtained for the pre-immune serum.
]]> RESULTS AND DISCUSSION
Experimental designs
Experimental designs allowed step by step sequential design of the process of purification of MY32/Ls protein for the API of a new vaccine against sea lice. In all cases the process parameters and conditions were established to obtain an API that meets the necessary requirements of a veterinary vaccine. Also the influence that the purification process could have on the IgG titers of the API regarding the analytical standard used was determined.
Establishment of cell rupture
The biomass obtained by the method described in the previous section yielded a 45-50 % of expression of MY32/Ls. A similar performance was obtained in the release of total proteins from passes 8 and 10 of mechanical rupture and the chemical rupture (7.94 g/L, 8.16 g/L and 7.78 g/L, respectively). The main disadvantage in the chemical rupture was the increased viscosity of the supernatant, which may have influenced on subsequent chromatographic steps. These rheological characteristics must be considered for the establishment of subsequent purification steps. The results corresponded with those obtained by Falconer et al. [10]. Chemical rupture had a 20 % lower cost per gram of protein of interest than the mechanical rupture and it is more attractive technologically. An analysis of the advantages and disadvantages of the methods used led to the selection of chemical rupture with 8 mol/L of urea as the most appropriate method, since it allows cell rupture and solubilization of inclusion bodies for MY32/Ls protein extraction.
Establishment of the washing and elution conditions
It was determined that for each milliliter of CSFF matrix between 2.1 and 2.5 mg of MY32/Ls protein adhered at all pH values tested (Figure 1). For further studies, a load capacity of 2 mg of MY32/Ls/mL of matrix was assumed.
The assessment of the influence of pH on washing and elution (Figure 2) evidenced that pH changes did not removed completely the specific proteins adhered to the matrix. Hence, pH changes were inefficient for washing and elution of the MY32/Ls, considering that the last washing/elution step (elution B) using 0.5 mol/L imidazole was able to extract the protein still bound to the matrix.
]]> The main disadvantage of this method is the handling of high working volumes in the washing and elution steps of the process, making it technologically unattractive. Above 80% presence of the representative MY32/Ls protein band was observed in the elutions, with a 73.8 % of total MY32/Ls protein recovery in the process, of 49.2 % in the elution step.The use of different imidazole concentrations was evaluated to establish the washing and elution steps, their results shown in figure 3 as determined by SDS-PAGE. In lane 9, corresponding to 0.04 mol/L of imidazole, the presence of the band of interest is not observed. Nevertheless, 0.1 mol/L imidazole was chosen in the washing step to guarantee an effective washing, and 0.5 mol/L in the elution step since a significant protein band was still present at that concentration. Protein MY32/Ls, unlike other proteins [11], elutes at high imidazole concentrations, what increase the costs due to the consumption of this raw material.
The defined imidazole concentrations were chromatographically tested and determined by SDS-PAGE (Figure 4). The presence of the representative MY32/Ls protein band in the elutions was observed above 80 %. The total protein recovery of MY32/Ls was 41 % and 37 % in the elution step.
Comparing the two methods for establishing the chromatographic step, it was determined that the total costs are reduced by 12.33 % with the use of imidazole for washing and elution, compared to washing and elution by pH changes. This reduction in costs is influenced mainly by the raw materials and reagents.
Scaling the chromatographic step to pilot scale
Two chromatographic runs were made and they were further evaluated by SDS-PAGE (Figure 5). The samples corresponding to the elution showed distributions similar to that of the positive control, with a purity greater than 80 % for the major band as assessed by densitometry.
The operation method for this step was established as follows. Equilibration step: 1.8 column volumes (50 cm/h). Pass: volumes never exceeding 1 column volume (10 to 12 cm/h) as limited by values obtained in rupture. First pass pushing step: 0.2 column volumes of equilibration buffer. Washing: 2 column volumes of washing buffer (25 cm/h). Elution: 1.5 column volumes of elution buffer (25 cm/h). Second pass pushing step: 0.5 column volume of water. Washing is sampled by collecting the two volumes of washing buffer and the first 0.5 volumes of Elution. The operation method is detailed in the table, with relevant description including the total proteins concentration and purity of MY32/Ls in each step.
]]> The three chromatographic runs of the Economic variant are shown in figure 6. The values of total proteins concentration were found in the aforementioned range (Figure 5) during the establishment of this operation method.Evaluation of the renaturation and dialysis step
The fact that Tween 20 and Tween 80 are not dialyzed by ultrafiltration membranes at concentrations higher than their critical micelle concentration (CMCTween 20 = ~0.00007 mol/L and CMCTween 80 = ~0.000012 mol/L) [12] was considered for the concentration and dialysis steps. As shown in figure 7, the influence of Tween 20 and 80 was similar; only at amounts above 1.2 mg per mg of total proteins does a significant difference start to appear.
A search for the maximum allowable concentrations to inoculate both additives intramuscularly was conducted. The maximum values for Tween 20 (0.0292 %) and Tween 80 (12 %) were established [13]. Therefore, we decided to use Tween 80 at 2 mg per mg of total protein as additive in the renaturation/dialysis step. An API was obtained showing bands of molecular weights higher than that of protein monomers, but recognizable by Western Blot (Figure 8). Such higher weight bands could be the subject of further adjustments in this step.
Evaluation of MY32/Ls immunogenicity
The IgG antibody response obtained in the three groups of C57Bl/6 mice demonstrated that the API obtained according to the purification process proposed was more immunogenic than the MY32/Ls protein standard (titers 2.7 times higher on day 28 and 3.5 times higher on day 58) (Figure 9). This may indicate that the API purification process effectively eliminates trace compounds present in the experimental MY32/Ls standard preparation which interfere with an adequate immunological recognition of the protein.
]]>CONCLUSIONS
The development of compact protein purification processes is an effective alternative to reduce production costs of new active APIs for veterinary use. In this sense, the chemical rupture with 8 mol/L urea is a more efficient alternative technically and economically than rupture with glass beads for the MY32/Ls protein. In this step, the protein of interest was obtained in soluble form and with the necessary conditions for use in metal ion affinity chromatography. On the other hand, in the chromatographic step, it was necessary to ensure the presence of imidazole at concentrations of 0.1 mol/L and 0.5 mol/L, respectively for washing and elution of MY32/Ls. This protein required to be stabilized with 2 mg of Tween 80 per mg of total proteins. The designed process provides an MY32/Ls API able to induce higher antibody titers than those generated by the research-scale protein standard tested.
REFERENCES
1. Pike AW, Wadsworth SL. Sea lice on salmonids: their biology and control. Adv Parasitol. 1999;44:233-7.
]]>2. Ragias V. Incidence of an intense Caligus minimus Otto 1821, C. pageti Russel, 1925, C. mugilis Brian, 1935 and C. apodus Brian, 1924 infection in lagoon cultured sea bass (Dicentrarchus labrax L.) in Greece. Aquaculture. 2004;242:727-33.
3. Johnson SC, Fast MD. Interactions between sea lice and their hosts. Symp Soc Exp Biol. 2004(55):131-59; discussion 243-5.
4. Carpio Y, Garcia C, Pons T, Haussmann D, Rodriguez-Ramos T, Basabe L, et al. Akirins in sea lice: first steps towards a deeper understanding. Exp Parasitol. 2013;135(2):188-99.
5. Carpio Y, Basabe L, Acosta J, Rodriguez A, Mendoza A, Lisperger A, et al. Novel gene isolated from Caligus roger-cresseyi: a promising target for vaccine development against sea lice. Vaccine. 2011;29(15):2810-20.
]]> 6. Carpio Y, Estrada MP, inventors; Centro de Ingeniería Genética y Biotecnología, assignee. Amino acid and nucleic acid sequences and vaccine to control ectoparasite infestations in fish. International Patent WO 2008145074 A3; 2009 Jan 29.7. CSFF - IMAC. Fairfield: General Electric; 2012.
8. Weber K, Osborn M. The reliability of molecular weight determinations by dodecyl sulfate-polyacrylamide gel electrophoresis. J Biol Chem. 1969;244(16):4406-12.
9. Alegria-Schaffer A, Lodge A, Vattem K. Performing and optimizing Western Blots with an emphasis on chemiluminescent detection. Methods Enzymol. 2009;463: 573-99.
10. Falconer RJ, O’Neill BK, Middelberg AP. Chemical treatment of Escherichia coli: 1. Extraction of intracellular protein from uninduced cells. Biotechnol Bioeng. 1997; 53(5):453-8.
11. Vancan S, Alves E, Alves SM. IMAC of human IgG: studies with IDA-immobilized copper, nickel, zinc, and cobalt ions and different buffer systems. Process Biochem 2002;37(6):573-79.
12. Pierce Biotechnology. Tech Tip #19. Remove detergent from protein samples. TR0019.2. 2009 [cited 2013 Nov 14]. Available from: http://proteomicsresource.washington.edu/docs/protocols03/Thermo_TechTip19_Detergent_Removal.pdf
13. Inactive Ingredient Search for Approved Drug Products. 2012 [cited 2012 Jan 20]. Available from: http://www.access-data.fda.gov/scripts/cder/iig/index.cfm
Received in November, 2013.
Accepted in April, 2015.
Carlos E Perez-Heredia. Grupo de Desarrollo Tecnológico, Centro de Ingeniería Genética y Biotecnología (CIGB). Circunvalación Norte y Ave. Finlay. Camagüey, Cuba. E-mail: carlos.perez@cigb.edu.cu
]]>