My SciELO
Services on Demand
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Biotecnología Aplicada
On-line version ISSN 1027-2852
Biotecnol Apl vol.31 no.2 La Habana Apr.-June 2014
RESEARCH
In vitro modeling of the environmental performance of attenuated strains of Vibrio cholerae O139
Modelación in vitro del comportamiento ambiental de cepas atenuadas de Vibrio cholerae O139
Talena Ledón, Daneylis D Hernández, Karen Marrero, Rafael Fando
Departamento de Biología Molecular, Dirección de Enfermedades Infecciosas, Centro Nacional de Investigaciones Científicas, CNIC. Ave. 25 y 158, CP 10600, Playa, La Habana, Cuba.
ABSTRACT
Vibrio cholerae, serogroups O1 and O139, is the causative agent of cholera diarrheal disease. Much of the research aimed to develop oral cholera vaccines is directed to the production of live attenuated strains, such as the V. cholerae O139 TLP01 and TLP05 strains. These two strains lack CTXφprophage genes and do not produce the hemagglutinin protease, a relevant pathogenesis factor, and the mannose-sensitive hemagglutinin fimbria, which could play an important role in the environmental behavior. In this work, different in vitro models were used to study the potential environmental performance of these two strains. Their ability to produce different types of biofilms, to acquire rugose phenotype and to resist different environmental stress conditions such as the presence of chlorine, detergents or high salt concentrations, were assessed. Significantly, the TLP01 and TLP05 strains displayed characteristics that limit their in vitro survival under different stress conditions, with respect to controls and wild-type strains. Such behavior under harsh environmental settings may limit their survival and act for their containment while using them as active ingredients for the development of a cholera vaccine candidate.
Keywords: cholera vaccine, Vibrio cholerae O139, attenuated strains, environmental persistence.
RESUMEN
El cólera es una enfermedad diarreica producida por la infección con la bacteria Vibrio cholerae de los serogrupos O1 y O139. Muchas de las investigaciones para el desarrollo de vacunas orales contra esta enfermedad se dedican a la producción de cepas vivas atenuadas. TLP01 y TLP05 son cepas atenuadas del serogrupo O139, que carecen de los genes del profago CTXφ; además, no producen la hemaglutinina proteasa, importante factor de la patogénesis; ni la fimbria hemaglutinina sensible a manosa, la cual pudiera ser importante en el comportamiento de las cepas vacunales en el ambiente. En este artículo se describe el posible comportamiento ambiental de estas cepas, a partir del estudio in vitro de sus propiedades para producir varios tipos de biopelículas, de adquirir el fenotipo rugoso y de resistir varias condiciones de estrés ambiental, como la presencia de hipoclorito de sodio, de detergentes o altas concentraciones salinas. Los resultados evidencian que las cepas TLP01 y TLP05 poseen características que limitan su supervivencia in vitro frente a distintas condiciones de estrés, con respecto a cepas controles y salvajes. En el entorno ambiental, estas características pudieran limitar su supervivencia y actuar como elementos de contención del agente biológico que se use como ingrediente farmacéutico activo del candidato vacunal.
Palabras clave: vacuna de cólera, Vibrio cholerae O139, cepas atenuadas, persistencia ambiental.
INTRODUCTION
Cholera is a diarrheic disease caused by the infection of the Vibrio cholerae bacterium. Symptoms are produced by its toxin, known as cholera toxin (CT), which is secreted in the small intestine. Only V. cholerae serogroups O1 and O139 display epidemic potential, with no crossprotection against both serogroups due to differences in their somatic components [1].
The disease is characterized by sudden outbreaks of fast spread, with health systems commonly collapsing when facing the outbreaks, particularly in underdeveloped countries. Prevention measures are recommended, such as guaranteeing drinking water sources and their permanent sanitation, real challenges under economic constraints [2]. Therefore, a successful vaccine has been considered advantageous for prevention. One of the leading strategies in the field of cholera vaccination is focused on obtaining a live-attenuated V. cholera strain, which could be administered by the oral route to stimulate the mucosal immune system in a fashion similar to that of wild-type bacterial strains. This could lead to the development of promising vaccine candidates able to mimic the events of the natural infection [3].
The use of that vaccine strategy implies concerns on the reacquisition by engineered strains of toxin genes from wild-type strains. In fact, the CT coding genes are carried by the CTXφfilamentous phage genome [4]. In this sense, the probabilities for virulence reversion by phage infection would be limited mainly to conditions promoting bacterial expression of the phage receptor, mainly at the intestines. There are other vibriophages: VGJφ, VEJφ, KSF-φ, VSK, VSKK and fs2, which could display advantages over CTXφto transmit the CT coding genes between strains in the natural ecosystems [5-7]. These phages use the mannose-sensitive hemagglutinin fimbria, which is expressed in watery environments and may be involved in the transmission of the toxin genes by hybrid CTX phages through a specialized transduction mechanism.
In this line of development for vaccine purposes, two live-attenuated V. cholerae strains were recently obtained at the National Center for Scientific Research (CNIC), in Cuba, based on serogroup O139 and named TLP01 and TLP05. They were generated by deletion of CTXφ prophage and CT ctxAB genes, followed by the replacement of the hapA gene encoding for the protease hemagglutinin by an inactivated version. This last bears an inserted DNA fragment coding for the endoglucanase A from the bacterium Clostridium thermocellum, used for easy selection of the engineered strains. Additionally, the mshA gene was replaced by a mutated allele to prevent the production of the MSHA fimbria. This aborts the infection by the CT gene-carrying hybrid phages using fimbria as receptor and reduces the probabilities for CT genes reacquisition through this mechanism [8].
As evidenced so far, the incidence of cholera outbreaks is closely related to environmental and ecological factors which are subsequently controlled by large-scale climate changes [9, 10]. Microorganism persistence fundamentally depends on its ability to adapt and develop survival strategies, among them: biofilm formation, rugose phenotype acquisition or reaching the viable non-culturable state [11]. In fact, V. cholerae commonly grow forming biofilms on either biotic or abiotic surfaces [12]. Those biofilms provide a microenvironment probably favoring microorganism survival and persistence for long interepidemic periods, by supporting the establishment of positive metabolic transactions with other bacterial community members. Besides, biofilms confer protection against several stressing and predatory environmental agents [13].
V. cholerae biofilm formation depends on the synthesis of an exopolysaccharide (EPS) coded by a set of genes located in the vps polysaccharide operon. Strains O1 El Tor and O139 additionally depend on flagellar motility for biofilm production, and O1 El Tor strains further require the production of the MSHA fimbria [14, 15]. The acquisition of the rugose phenotype is also included among the attributes needed to produce biofilms [16].
There are also reports on V. cholerae development of other types of biofilms, independent of vps genes, in a seawater model. They required Ca2+ ions instead of the vps genes monosacharides [17].
However, if the abovementioned TLP01 and TLP05 strains of V. cholerae serogroup O139 would be used as active pharmaceutical ingredients for vaccine development, it is expected that vaccinees will excrete these strains in the feces. Therefore, it would be necessary to evaluate or to model the environmental impact of their release.
For that purpose, this work was aimed to model the environmental performance of the V. cholerae TLP01 and TLP05 vaccine strains. Their abilities to produce different types of biofilms, to acquire the rugose phenotype and persist under stressing environmental conditions (sodium hypochlorite, detergents or high salt concentrations) were evaluated in vitro.
MATERIALS AND METHODS
Materials
Strains
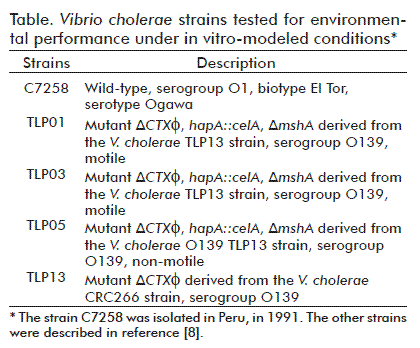
The V. cholerae strains used in this study were: C7258, wild-type; TLP01 and TLP05, mutant strains to the mshA gene; TLP03, an isogenic strain; and TLP13 atoxigenic parental strain (Table). The last two strains were used as assay controls.
Culture media
The culture media were TSB (17 g/L tryptone, 3 g/L peptone, 2.5 g/L K2HPO4, 5 g/L NaCl) supplemented at 0.4 % glucose; simulated seawater (27.3 g/L NaCl, 13.6 g/L Mg2SO4, 1.5 g/L CaCl2, 0.8 g/L KCl, 0.3 g/L NaHCO3, 0.005 g/L Na2B4O7, 0.001 g/L LiCl), adjusted to pH 7.0 and supplemented with casein hydrolysate at 1 %; and peptone alkaline water (10 g/L peptone, 10 g/L NaCl; pH 8.5).
Methods
Assessment of biofilm formation ability
The ability of V. cholera strains to produce biofilms was assessed in TSB medium supplemented with glucose and simulated seawater [17]. The assay was conducted in 96-well polystyrene plates, starting from a 1:100 dilution of fresh cultures at exponential phase with similar optical densities at 600 nm (OD600nm). This avoided the presence of autoinducers in the inoculums which may interfere in biofilm formation. Wells containing uninoculated culture media were used as baseline controls [18]. The strains TLP01, TLP03, TLP05 and TLP13 were tested, and the experiments were run in triplicate. The plates were incubated at 37 ºC for 24, 48 or 72 h, and culture homogeneity was checked at each time point by determining OD600nm in a microplate reader (Multiscan, Finland). After three successive washes in phosphate-buffered saline (PBS; 137 mM NaCl, 9.58 mM Na2HPO4, 2.68 mM KCl, 1.47 mM KH2PO4; pH 7.2), plates were stained with 1 % safranin solution (prepared in 33 % acetic acid), and biofilm OD490nm of biofilms was measured in the same microplate reader.
Induction of the rugose phenotype
Alkaline peptone water 5-mL samples were inoculated with 5 µL of fresh cultures of strains TLP01, TLP05 or TLP13, and incubated at 37 ºC for 30 days. Serial dilutions were made on preset time points and strains were plated in LB-agar medium (10 g/L tryptone; 10 g/L NaCl; 5 g/L yeast extract, pH 7.0, plus 15 g/L bacto agar). Strain morphology was observed and classified as rugose or flat, also describing any other variation if present [19].
Assessment of strain sensitivity to stressing agents
The sensitivity of strains TLP01, TLP05 and TLP13 was tested under stressing environmental conditions, comprising the presence of sodium hypochlorite, sodium dodecyl sulfate (SDS) and high salt concentration.
Sensitivity to sodium hypochlorite
The survival of V. cholerae strains TLP01 and TLP13 to sodium hypochlorite concentrations was tested under conditions favoring or not biofilm formation [19]. First, strains were grown in capped borosilicate flasks filled with 3 mL of TSB medium supplemented at 0.4 % glucose, and incubated at 30 ºC for 24 h. Subsequently, planktonic cultures of each strain were checked for homogeneity in a spectrophotometer for OD630nm. The planktonic content was discarded and aggregated cells were resuspended in 5 mL of PBS. Biofilms were disaggregated by adding 1 g of glass pearls to each flask followed by vortexing. Samples were taken for microcounting by serial dilutions, prior to the addition of the stressing agent. Then, NaOCl was added up to 2 ppm and incubated for 1, 5, 10, 20 or 30 min. Subsequently, the stressing agent was inactivated on the given samples by adding sodium thiosulfate at 0.015 %, viable cells were counted, and adequate amounts of each dilution plated in LB and further incubated at 37 ºC for 20-22 h.
Strains were also cultured under shaking to avoid biofilm formation, in test tubes filled with TSB medium supplemented with glucose at 0.4 %, at 37 ºC for 4 h. Approximately 107 colony-forming units (c.f.u.) of each culture were subjected to the action of the previously mentioned NaOCl concentrations and their sensitivity to the agent was assessed as previously described.
Sensitivity to SDS
Strains TLP01, TLP05 and TLP13 were tested for sensitivity to sodium dodecyl sulfate (SDS), starting from cultures in a 24-well polystyrene plate containing TSB medium supplemented with glucose at 0.4 %, which were inoculated from 1:100 dilutions. A well containing uninoculated culture medium was used as baseline control. SDS 0.05, 0.1, 0.25, 0.5 and 1 % concentrations were tested [20]. The growth of each strain in the absence of the agent was also tested. The plate was read every 2 h at 630 nm. The experiment was done in duplicate.
Sensitivity to NaCl concentrations
For strains C7258, TLP01, TLP05 and TLP13, two colonies were isolated and TLP13, and further grown with agitation for 6 h. Afterwards, 1 mL of each bacterial culture was centrifuged and cells were resuspended in 1 mL of TSB medium supplemented at 2.5 M NaCl. After incubation for 0, 15, 30 and 60 min, 100 µL were taken for microcounting from each test sample. Serial dilutions were made from each and plated on LB medium, being further incubated at 37 ºC for 20-22 h. The results were analyzed by adjusting the fractions of bacteria surviving to salt stress to a curve, according to the Weibull model (a statistic distribution model of inactivation time points) [21]:
Where:
S: fraction of viable bacteria at each time point, in respect to the amount at the start of the experiment; calculated following the expression:
t: the period that the system is under the stress tested (salt stress);
α: scale parameter; depends on the evaluated component (osmolarity);
β: shape parameter; related to the curve shape and the resistance ability to the stress applied (salt stress).
Statistical analysis
Data were statistically analyzed with the Prisma for Windows statistical package, version 4.0. The statistical significance was set to 0.05 % for all the comparisons. Means were compared by the Student’s t test as indicated for each case in the Results and discussion section.
RESULTS AND DISCUSSION
Biofilm formation ability
The ability of V. cholerae TLP strains to form biofilms was assessed, under conditions promoting them or not depending on EPS production (vps-dependent or vps-independent biofilms). Studies were run in rich medium and simulated seawater, respectively. The ability was inferred from the quantification of the EPS produced by each strain, due to the tight relationship between these processes [14, 15].
In the absence of EPS, bacteria only formed a monolayer on the culture surface [14]. Similarly, in the presence of any other mutation blocking biofilm formation, the strain produces undetectable EPS amounts, according to the test method used.
Quantification of vps-dependent biofilms
V. cholerae forms dense biofilms composed of bacterial columns surrounded by water channels in rich culture media such as LB or TSB [15]. These media provide the ideal conditions in vitro to promote vps-dependent biofilm production.
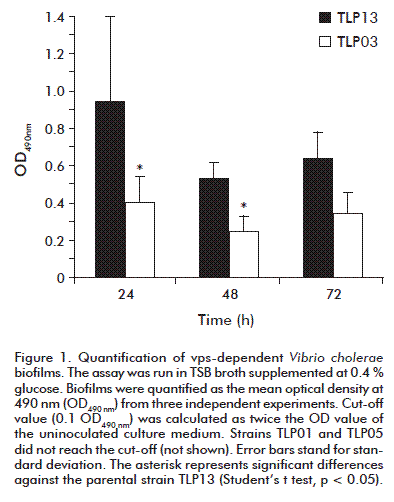
In TSB medium supplemented with glucose, only the TLP03 and TLP13 strains produced detectable EPS amounts within the first 24 h, supporting the assumption that they do produce biofilms. A halo was detected in the air-liquid interphase, characteristic of those tridimensional structures. Nevertheless, strains TLP01 and TLP05 did not produced detectable EPS (Figure 1). Such evidences indicate that, even when strains TLP03 and TLP13 generate biofilms under the conditions tested, this ability was significantly reduced for TLP03 (an mshA mutant) compared to the atoxigenic parental strain TLP13, at least after 24 and 48 h of culture (Student’s t test, p = 0.018 and p = 0.0134, respectively). The absence of biofilms for the TLP05 strain could be related to its non-motile phenotype. This strain carries a random mutation unrelated to its engineering process which abrogates motility, although its flagellum was present as corroborated by electron microscopy [8]. Flagellar movement influences biofilm formation in V. cholerae O139, as reported by Watnick et al. [15]. This process is particularly relevant at starting the monolayer.
The results indicated that a spontaneous mutation had to occur in the TLP01 mutant interfering on the formation of these structures. It is unrelated to the mshA mutation, since the evidences suggest that the isogenic TLP03 mutant could establish the tridimensional polysaccharide structure.
Quantification of vps-independent biofilms
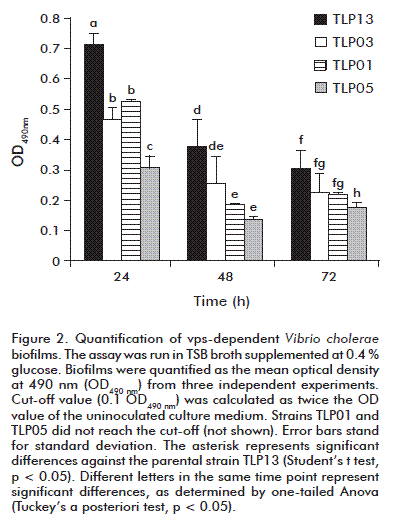
In seawater, V. cholerae forms calcium-dependent biofilms instead on vps genes. This element directly mediates cell-to-cell or cell-surface interactions in bacterial biofilms [17]. Hence, a seawater model obtained in the laboratory [17] was chosen to quantify vps-independent biofilm production. As established from readouts at 600 nm, strains TLP01, TLP03, TLP05 and TLP13 similarly produced biofilms under the tested culture conditions (Figure 2).
In contrast to TSB medium results, all the strains were able to produce biofilms in seawater. There are reports describing that MSHA is required but unessential for the calcium-dependent biofilm formation and that mshA mutants establish weaker cell-to-cell interactions and thinner biofilms [17]. Our results are in agreement with those reported, since mshA mutants showed reduced biofilm formation compared to the parental TLP13 strain (Figure 2).
Regarding motility, under sugar deficitary conditions and in the presence of calcium, the flagellum could become a signal inducing biofilm formation. According to Kierek and Watnick [17], the flagellum is also required to structure calcium-dependent structures, but with an effect not as relevant as in the case of the equivalent mutation for vps-dependent biofilms. The results obtained for the TLP05 strain also confirms those previous reports. During the experiment, TLP05 generated biofilms at levels below those of the TLP01 and TLP03 mutants, at least after 24 h (One-tailed Anova, p < 0.05; Tuckey’s a posteriori test, p < 0.05). Hence, the TLP05 strain could be less prone to produce biofilms due to the mshA gene mutation and its non-motile phenotype.
These results indicate that TLP01 and TLP05 strains produce vps-independent biofilms; though, at levels below those of the parental strain. Accordingly, under the studied conditions, these two mutants do not bear any phenotype-related competitive advantage for environmental persistence. This type of biofilms requires millimolar calcium concentrations, absent in fluvial environments but found in seawater [22]. Such calcium dependence determines that vps-independent biofilms could be more relevant in marine ecosystems than in lacustrine and briny ones.
On the other hand, biofilm formation is required to reach the viable non-culturable state [11]. In this sense, the inability of the TLP01 and TLP05 mutants to produce biofilms could lead to their decreased survival and make them disadvantageous in natural environments which are far more complex than conditions modeled in this study. Thus, a different behavior cannot be ruled out in natural environments, although such non-biofilm producer phenotype could be a containment element contributing to the biosafety of the live-attenuated vaccines having those strains as active pharmaceutical ingredients. Other experiments must be run, including competition assays with wild strains, in order to evaluate other types of interactions in a scenario closer to the natural environments.
Induction of the rugose phenotype
The V. cholera rugose phenotype is a bacterial survival and environmental persistence mechanism. This morphological change makes it highly resistant to agents as chlorine, hydrogen peroxide and other oxidative or osmotic stresses [19]. Previous studies demonstrated that the rugose phenotype is related to EPS production, which promotes biofilms structuring [16]. Keeping this in mind, the development of rugose phenotype was evaluated for TLP01 and TLP05 mutants compared to the TLP13 parental strain, by incubation in alkaline peptone water at 37 ºC for 30 days. It was shown that the atoxigenic parental strain TLP13 started the rugose phenotype on day 8. Meanwhile, from day 13 on, the TLP01 strain developed opaque colonies with a corolla in the center but not the typical rugose phenotype. The TLP05 mutant kept the flat phenotype during the entire experiment.
These results confirmed that TLP01 and TLP05 had lost the ability to develop rugose colonies and, therefore, biofilms in vitro, this ability intact in its parental TLP13 strain. These two phenotypes depend on EPS production, indicating that TLP01 and TLP05 mutants have their vps-dependent EPS production mechanism affected at certain point not engineered during their genetic construction. Such a dysfunction would explain both, their inability to develop the rugose phenotype and the tridimensional structuring of biofilms.
Those properties would make TLP01 and TLP05 more vulnerable to environmental stressing conditions, affecting their persistence in aquatic ecosystems where nutrients are far restricted and resistance mechanisms required. Nevertheless, the relevance of EPS for environmental persistence is debatable since there are no reports on rugose isolates from environmental niches.
Assessment of strain sensitivity to stressing agents
Sensitivity to chloride
Sodium hypochlorite is a first-choice disinfectant for drinking water regardless its instability, being used at concentrations ranging 2-5 ppm. It has been reported to induce the rugose phenotype in V. cholerae, as for other agents. This also supports the environmental disadvantage of TLP01 and TLP05 compared to other natural strains. At the same time, biofilm formation arises as another resistance mechanism, with the agent inactivated after getting into contact with polysaccharide structures. Therefore, resistance to chloride was studied in the TLP01 and TLP13 strains which produce or not biofilms. The experiment was run under conditions favoring or not biofilm formation (planktonic or shaked cultures, respectively), by exposure of 106-107 c.f.u./mL to 2 ppm for 1, 5, 10, 20 and 30 min, and surviving c.f.u. were counted.
Strains grown under shaking had increased chloride sensitivity (Figure 3A), with no viable cells after 5 min of exposure. It has been described that chloride is a stress agent for V. cholerae [19]. The planktonic culture characterized by free cells in the medium provides no protection against the agent, as evidenced in the experiment. On the contrary, the atoxigenic parental strain TLP13 was highly resistant when biofilms were present, with steady viable cell counts over time (Figure 3B). There were about 107 c.f.u./mL after a 30-min exposure, opposed to the lack of viable cells at any time for the TLP01 strain (Figure 3B). These results were in agreement with those of cultures under shaking.
Our findings confirm the relevance of TLP13 biofilms in chloride inactivation and allow to foresee the relevance these structures would play in V. cholerae survival and persistence. In spite of the environmental disadvantage of strains TLP01 and TLP05, their coexistence with other strains able to produce such structures could mitigate their limitations.
Sensitivity to SDS
The sensitivity of V. cholerae to SDS at concentrations ranging 0.05-1.0 % was investigated. The typical performance of strains TLP01, TLP05 and TLP13 in TSB medium supplemented with glucose at 0.4 % in the absence of SDS was set as reference condition for the experiment (Figure 4A). There were no differences in the growth rate among the three strains, as evidenced by spectrophotometric determinations at 630 nm.
When SDS was tested, at 0.05 % cultures grew similarly for all the strains, with no significant delays. The stressing effect of SDS manifested at 0.1 % concentrations although growth was not completely inhibited. The growth of mutant strains TLP01 and TLP05 stopped at 0.25 %, with strain TLP13 still growing under those conditions and only stopped at 1.0 % SDS (Figure 4B). These results indicate that the SDS minimal inhibitory concentration is four times lower for TLP01 and TLP05 than that of the parental TLP13.
As most of gram-negative bacteria do, V. chole-rae regulates the diffusion rate of small molecules through its external membrane by modulating the synthesis of external membrane porins, specifically OmpT and OmpU [23]. In response to bile salts, the transcriptional regulator ToxR positively induces the expression of OmpU, while suppressing that of ompT. The production of OmpT is associated to the increase in bacterial bile sensitivity [20, 24], while the OmpU production has been implicated in resistance to antimicrobial peptides, organic acids and anionic bile detergents and SDS [25]. Altogether, these results allow predicting the performance of the strains tested, in the presence of other agents of similar nature such as antibiotics and other types of detergents.
It has been previously described a major OmpU band in the SDS-PAGE electrophoretic pattern of external membrane proteins of the TLP13 wild type strain [26]. This pattern is characteristic of strains carrying the toxR wild-type allele. On the contrary, the toxR mutants only produce OmpT. Curiously, the TLP01 and TLP05 mutants show detectable levels of both proteins, indicative of an intermediate phenotype probably caused by a mutation [26]. According to the results of Provenzano and Klose [24], the reduced resistance to detergents as SDS in these two mutant strains may be related to a lower expression of the OmpU porin. Otherwise, the effect that other mutations could exert in these vaccine strains cannot be ruled out.
Osmotolerance
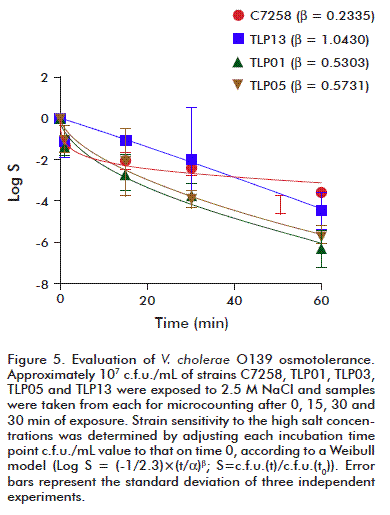
High salinity is among the harsh physicochemical conditions that could affect the environmental performance of V. cholerae [27]. Hence, a test was run to evaluate the effect of high NaCl concentrations on the outcome of the TLP01, TLP05 and TLP13 strains. The wild-type C7258 strain was used as control. Approximately 107 c.f.u./mL of each strain was subjected to the effect of the stressing agent. Results were analyzed by adjusting bacterial fractions, which survived after 15, 30 or 60 min of exposure to NaCl, to a curve according to the Weibull model. That model has been successfully used to evaluate tolerance of different microorganism’s populations against stressing agents [21]. Results are summarized in figure 5. Viable cell counts declined over time for all the strains, with TLP01 and TLP05 c.f.u. counts two orders below those of the wild strain, very significant for the study.
The β constant values for each strain curves were compared by the Fisher’s test. As shown by comparing the control strain C7258 results to those of the other strains, it displayed a higher resistance to salinity (p < 0.05). The concave shape of its curve, influenced by its β value, could allow us to infer that C7258 cells have the ability to adapt to high salt concentrations, an effect absent for the other strains tested. There were no significant differences in the performance of the atoxigenic TLP01, TLP05 and TLP13 strains.
These results point towards osmoadaptation properties of the candidate vaccine strains, compared to a non-engineered strain. In this sense, TLP01 and TLP05 appear to be less tolerant to salt stress than the wild-type strain C7258. This is relevant considering than once released to the environment, they would be disfavored in respect to the natural toxigenic strains. At the same time, their reduced osmotolerance equally contributes to its biosafety when used as live-attenuated vaccines. Since there were no significant differences in the performance of TLP01 and TLP05 compared to TLP13 under the assayed conditions, this could indicate an intrinsic property of the two strains, or be related to a single collateral unexpected mutation, which appeared during the initial CTXφ genes deletion event. In spite of being fortuitous, the resulting effect provides a positive property to these mutants which may contribute to limit their environmental survival.
Noteworthy, V. cholerae is one of the few Vibrio species successfully performing under low osmolarity conditions, always in the presence of sodium, in spite of its optimal salinity ranging 5-25 g/L. It could also grow in dissolved organic material at salinity concentrations near 45 g/L. Therefore, this bacterium adapts well to freshwater and briny waters in endemic areas. The periodical intrusion of salt water seems to increase Vibrio survival in those environments, but in certain areas and for given periods the increase in salinity tends to reduce the viable cell counts [9]. In this sense, the strains acquiring the viable non-culturable state should be also considered. This last aspect remains to be evaluated in vaccine strains.
CONCLUSIONS
Here we demonstrate that strains TLP01 and TLP05 had limitations in the mechanisms that regulate biofilm and EPS formation in vitro, osmotolerance and susceptibility to detergents. The impact of these properties under other experimental conditions near those of the natural ecosystems should be further studied, in spite of these mechanisms being expected to influence the vibrios environmental performance to certain extent. The results indicate that these two strains have properties which limit their behavior in vitro. And such properties could act as containment for the biological agent used as active pharmaceutical ingredient for a vaccine candidate.
The mshA gene mutation also reduces the possibility to reacquire toxin genes in watery environments. The potential environmental persistence limitations coming from the results presented will contribute to increase vaccine biosafety. In this sense, a poor environmental performance could reduce the impact that the reacquisition of the toxin genes would have during human infection or for the environment. It seems to be quite improbable that these two strains could achieve bacterial population levels under competitive scenarios, enough to significantly contribute to disseminate the ctxAB genes or to affect humans.
REFERENCES
1. Faruque SM, Chowdhury N, Kamruzzaman M, Ahmad QS, Faruque AS, Salam MA, et al. Reemergence of epidemic Vibrio cholerae O139, Bangladesh. Emerg Infect Dis. 2003;9(9):1116-22.
2. Bishop AL, Camilli A. Vibrio cholerae: lessons for mucosal vaccine design. Expert Rev Vaccines. 2011;10(1):79-94.
3. Lopez AL, Clemens JD, Deen J, Jodar L. Cholera vaccines for the developing world. Hum Vaccin. 2008;4(2):165-9.
4. Waldor MK, Mekalanos JJ. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science. 1996;272(5270):1910-4.
5. Campos J, Martinez E, Marrero K, Silva Y, Rodriguez BL, Suzarte E, et al. Novel type of specialized transduction for CTX phi or its satellite phage RS1 mediated by filamentous phage VGJ phi in Vibrio cholerae. J Bacteriol. 2003;185(24):7231-40.
6. Campos J, Martinez E, Izquierdo Y, Fando R. VEJphi, a novel filamentous phage of Vibrio cholerae able to transduce the cholera toxin genes. Microbiology. 2010;156(Pt 1):108-15.
7. Das B, Bischerour J, Barre FX. Molecular mechanism of acquisition of the cholera toxin genes. Indian J Med Res. 2011;133:195-200.
8. Ledon T, Ferran B, Perez C, Suzarte E, Vichi J, Marrero K, et al. TLP01, an mshA mutant of Vibrio cholerae O139 as vaccine candidate against cholera. Microbes Infect. 2012;14(11):968-78.
9. Lipp EK, Huq A, Colwell RR. Effects of global climate on infectious disease: the cholera model. Clin Microbiol Rev. 2002;15(4):757-70.
10. Huq A, Sack RB, Nizam A, Longini IM, Nair GB, Ali A, et al. Critical factors influencing the occurrence of Vibrio cholerae in the environment of Bangladesh. Appl Environ Microbiol. 2005;71(8):4645-54.
11. Kamruzzaman M, Udden SM, Cameron DE, Calderwood SB, Nair GB, Mekalanos JJ, et al. Quorum-regulated biofilms enhance the development of conditionally viable, environmental Vibrio cholerae. Proc Natl Acad Sci USA. 2010;107(4):1588-93.
12. Chiavelli DA, Marsh JW, Taylor RK. The mannose-sensitive hemagglutinin of Vibrio cholerae promotes adherence to zooplankton. Appl Environ Microbiol. 2001;67(7):3220-5.
13. Reidl J, Klose KE. Vibrio cholerae and cholera: out of the water and into the host. FEMS Microbiol Rev. 2002;26(2):125-39.
14. Watnick PI, Kolter R. Steps in the development of a Vibrio cholerae El Tor biofilm. Mol Microbiol. 1999;34(3):586-95.
15. Watnick PI, Lauriano CM, Klose KE, Croal L, Kolter R. The absence of a flagellum leads to altered colony morphology, biofilm development and virulence in Vibrio cholerae O139. Mol Microbiol. 2001;39(2):223-35.
16. Yildiz FH, Schoolnik GK. Vibrio cholerae O1 El Tor: identification of a gene cluster required for the rugose colony type, exopolysaccharide production, chlorine resistance, and biofilm formation. Proc Natl Acad Sci USA. 1999;96(7):4028-33.
17. Kierek K, Watnick PI. The Vibrio cholerae O139 O-antigen polysaccharide is essential for Ca2+-dependent biofilm development in sea water. Proc Natl Acad Sci USA. 2003;100(24):14357-62.
18. Yildiz FH, Dolganov NA, Schoolnik GK. VpsR, a member of the response regulators of the two-component regulatory systems, is required for expression of vps biosynthesis genes and EPS(ETr)-associated phenotypes in Vibrio cholerae O1 El Tor. J Bacteriol. 2001;183(5):1716-26.
19. Wai SN, Mizunoe Y, Takade A, Kawabata SI, Yoshida SI. Vibrio cholerae O1 strain TSI-4 produces the exopolysaccharide materials that determine colony morphology, stress resistance, and biofilm formation. Appl Environ Microbiol. 1998;64(10):3648-55.
20. Provenzano D, Klose KE. Altered expression of the ToxR-regulated porins OmpU and OmpT diminishes Vibrio cholerae bile resistance, virulence factor expression, and intestinal colonization. Proc Natl Acad Sci USA. 2000;97(18):10220-4.
21. Van Boekel MAJS. On the use of the Weibull model to describe thermal inactivation of microbial vegetative cells. J Food Microbiol. 2002;74(1-2):139-59.
22. Lizarraga-Partida ML, Mendez-Gomez E, Rivas-Montano AM, Vargas-Hernandez E, Portillo-Lopez A, Gonzalez-Ramirez AR, et al. Association of Vibrio cholerae with plankton in coastal areas of Mexico. Environ Microbiol. 2009;11(1):201-8.
23. Sperandio V, Bailey C, Giron JA, DiRita VJ, Silveira WD, Vettore AL, et al. Cloning and characterization of the gene encoding the OmpU outer membrane protein of Vibrio cholerae. Infect Immun. 1996;64(12):5406-9.
24. Provenzano D, Lauriano CM, Klose KE. Characterization of the role of the ToxR-modulated outer membrane porins OmpU and OmpT in Vibrio cholerae virulence. J Bacteriol. 2001;183(12):3652-62.
25. Bina XR, Provenzano D, Nguyen N, Bina JE. Vibrio cholerae RND family efflux systems are required for antimicrobial resistance, optimal virulence factor production, and colonization of the infant mouse small intestine. Infect Immun. 2008;76(8):3595-605.
26. Ledón T, Ferran B, Fando R. Estudio de la funcionalidad de ToxR en cepas atenuadas de Vibrio cholerae O139. Rev CENIC Cienc Biol. 2011;42(1):25-28.
27. Castaneda Chavez Mdel R, Pardio Sedas V, Orrantia Borunda E, Lango Reynoso F. Influence of water temperature and salinity on seasonal occurrences of Vibrio cholerae and enteric bacteria in oyster-producing areas of Veracruz, Mexico. Mar Pollut Bull. 2005;50(12):1641-8.
Received in September, 2013.
Accepted in April, 2014.
Talena Ledón. Departamento de Biología Molecular, Dirección de Enfermedades Infecciosas, Centro Nacional de Investigaciones Científicas, CNIC. Ave. 25 y 158, CP 10600, Playa, La Habana, Cuba. E-mail: talena.ledon@cnic.edu.cu.