My SciELO
Services on Demand
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO
Share
Ingeniería Electrónica, Automática y Comunicaciones
On-line version ISSN 1815-5928
EAC vol.39 no.2 La Habana May.-Aug. 2018
ORIGINAL ARTICLE
PyECG: A software tool for the analysis of the QT interval in the electrocardiogram
PyECG: Una herramienta de software para el análisis del intervalo QT en el electrocardiograma
Alexander A. Suárez LeónI, Carolina VaronII, Rik WillemsIII, Sabine Van HuffelII, Carlos R. Vázquez SeisdedosI
IUniversidad de Oriente, Santiago de Cuba, Cuba.
IIKU Leuven, Lovaina, Belgium.
IIIUniversity Hospitals Leuven, Lovaina, Belgium.
ABSTRACT
PyECG is a software tool for QT interval analysis in the electrocardiogram (ECG). The software is written in Python 3.6 and among its main features includes signal filtering, Q onset, R peak and T offset detection algorithms, classifiers for irregular heartbeat identification and rejection, and tools for easy correction of automatically generated annotations. Moreover, the software includes a signal quality assessment module in order to help the researcher deciding which lead should be used. The software tool computes and plots the QT Variability Index (QTVI) and QT dynamicity parameters. Since the software is designed for cardiologists and specialists with no or little programming skills, the graphical user interface is intuitive, compact and easy to use.
Keywords: software, ventricular repolarization, QT interval analysis, QTVI
RESUMEN
PyECG es una herramienta de software para el análisis del intervalo QT en el electrocardiograma (ECG). El programa está escrito en Python 3.6 y entre sus principales características se encuentran: filtrado de la señal, algoritmos de detección de puntos de inicio de la onda Q, pico de la onda R y final de la onda T, clasificadores para la identificación y eliminación de latidos irregulares así como herramientas para facilitar la corrección de las anotaciones generadas automáticamente. Además, el software incluye un módulo para la evaluación de la calidad de la señal con el objetivo de asistir al usuario en decidir la derivación óptima para el análisis. La herramienta permite calcular y graficar el índice de variabilidad del QT (QTVI) y los parámetros del análisis dinámico del QT. El software está diseñado para cardiólogos y especialistas con independencia a sus habilidades de programación por lo que la interfaz gráfica de usuario es muy sencilla, compacta e intuitiva.
Palabras claves: software, repolarización ventricular, análisis intervalo QT, QTVI
1. INTRODUCTION
Cardiovascular diseases (CVD) are the major cause of death in Cuba and worldwide. The electrocardiogram (ECG) is a cheap, well-known and noninvasive technique for the assessment of the electrical activity of the heart. Most of the diagnostic information in the ECG relies on the morphology of the waves and the duration of intervals such as RR and QT. RR interval variability has been thoroughly studied and its clinical significance has been established on diagnosis and prognosis of several cardiovascular and non-cardiovascular diseases [1-2].
The QT interval is associated with the ventricular repolarization of the heart. It starts at the beginning of the QRS complex and finishes at the end of the T-wave. Normal values of QT interval for males are around 450 ms, meanwhile, for healthy females, the mean value is 470 ms [3]. It is well known that QT interval is influenced by changes in heart rate and other factors like drug intake [4]. QT anomalies have been associated with the risk of suffering ventricular life-threatening arrhythmias and Sudden Cardiac Death (SCD). For instance, in both congenital and acquired Long QT Syndrome (LQTS) there is a risk of developing Torsade de Pointes (TdP), a kind of polymorphic ventricular tachycardia which may result into SCD [5]. Several markers have been proposed for the assessment of ventricular repolarization instability. The most relevant ones include QT variability indexes, QT dynamicity, QT dispersion (QTd) and T-Wave Alternans (TWA).
QT variability (QTV) is the quantification of the slight changes in the QT beat to beat. One of the most accepted QTV temporary indexes is the QTVI proposed by Berger et al[6], and it is defined as
where the terms QTV and HRV correspond to the variance of QT and RR segments, respectively, and QTM and HRM represent the average values of these segments in the range considered. QTVI has been studied in different populations, such as dilated cardiomyopathy (DCM) patients, post-Myocardial Infarction (MI) subjects, healthy controls, patients with implantable cardioverter-defibrillator devices (ICD) [7] and more recently, on patients that suffered spinal cord injury [8].
Different methods exist to assess repolarization dynamicity, also known as adaptation or hysteresis. A well-known method estimates QT/RR slope, i.e., the slope of the linear regression between QT and RR intervals. Steeper QT/RR may indicate either excessive lengthening of QT at slow rates or excessive shortening at fast rates. Both of these processes may contribute to the occurrence of a malignant ventricular arrhythmia and consequently to SCD. Although there is no global consensus on the normal values of QT/RR slope, small values are in general associated with healthy people whereas higher values correspond to pathological subjects [9].
Other relevant markers include QT dispersion (QTd) and T-Wave Alternans (TWA). QTd is defined as the difference between the maximum and minimum QT in a standard 12-lead ECG [10]. It mainly reflects regional variations in ventricular repolarization. However, there is still uncertainty on the ability of QTd to identify patients at high risk of sudden death [11]. Moreover, it requires a standard 12-lead ECG and is not available on routine used monitoring systems, but could be indirectly derived from Holter systems. TWA is a beat-to-beat alternation in the morphology and amplitude of the ST-segment or T-wave. It reflects spatiotemporal heterogeneity in the repolarization and has been proposed as a marker for risk stratification of ventricular tachycardia (VT) - ventricular fibrillation (VF) and SCD [12-13].
Several software tools have been developed for the analysis of the ECG signal, particularly for Heart Rate Variability (HRV) analysis. In [14], a software for advanced HRV analysis is presented. This tool is available free of charge on Linux and Windows platforms. gHRV is an open source tool written in Python for HRV analysis [15]. It is multiplatform software (Linux, Windows and Apple OS X) and provides support for several formats. Other works include a tool for HRV spectral analysis [16], a software for the analysis of cardio-respiratory variability signals [17] and a tool in Matlab® for the analysis of cardiac inter-beat interval (IBI) data [18]. An extended revision of software tools for HRV can be consulted in [19].
Although heartbeat classification in HRV software is rarely used, it is crucial in QT analysis because most of the indexes are defined for normal heartbeats. Hence, the inclusion of abnormal heartbeats, like premature atrial or ventricular beats, would bias the values of the indexes. Thus, a classification or labeling algorithm is needed for rejecting or identifying such beats. Ecg-kit is a package written in Matlab® scripting language which focuses on delineation and classification of the ECG signal [20]. However, this tool is intended to be Matlab® toolbox, so is not targeted to any specific task, which implies knowledge of the language or programming skills in order to exploit its features.
In spite of the considerable amount of studies in the analysis of ventricular repolarization, QT analysis is not always available in Holter systems and when it is present, the analysis is limited. There is also a lack of tools for computing QTVI using the methodology proposed by Berger [6]. Furthermore, up to the knowledge of the authors, there are no software tools that support research on QT analysis. The reason behind these facts could be that there is still a lot of research to do before including QT indexes in the standard diagnostic protocols. Nevertheless, as more research is needed there is also a need for tools that support such research. Hence, this paper proposes an open source software tool that uses advanced algorithms for the analysis of the QT interval in Holter recordings. The software called PyECG, is intended to be a useful tool for cardiologists and researchers with no or little programming skills, so a simple and intuitive graphical user interface is provided. The software is written in Python and supports both, QTVI analysis and QT dynamicity analysis. Future versions may include T-wave alternans and QTd analysis.
2. MATERIALS AND METHODS
The software has been developed in Python 3.6 using Anaconda 3-5.01 for Linux. The user interface was designed using PyQt5 which is a wrapper in Python of the multiplatform library Qt. Both standard packages, NumPy and SciPy were extensively used as well. NumPy is a base package for scientific computing with Python. It includes an efficient array object and useful functions for performing linear algebra operations, Fourier transform and random number generation. Scipy is a collection of algorithms for mathematics, science, and engineering based on Numpy. Graphics generation used PyQtGraph which is a graphics and user interface library for Python intended for use in data acquisition and analysis applications. Other packages such as Scikit-Tensor and Scikit-Learn were also used. Scikit-Tensor is a Python module for multi-linear algebra and tensor analysis whereas Scikit-Learn is a machine learning library for Python [21]. The development environment (IDE) was PyCharm 2017.2.4 from JetBrains. The general workflow of the software is shown in Figure 1.
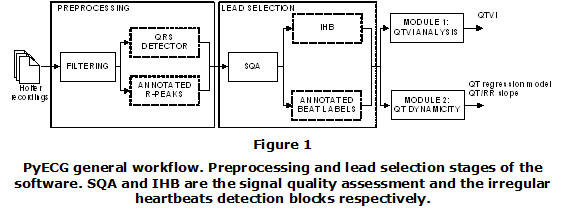
The tool consists of four main phases, see Figure 1. Since QT analysis requires a high-quality ECG signal some preprocessing steps are compulsory before doing any analysis. So, after loading the signal, the first two modules, namely Preprocessing and Lead selection, perform the signal conditioning, Figure 1. The preprocessing stage includes filtering and QRS complex detection whereas the lead selection stage includes a signal quality assessment (SQA) block and an optional irregular heartbeats detection block (IHB). Next, the user can select between two types of analysis. On one hand, Module 1 is intended for QTVI analysis in both, single and successive 256-second windows. On the other hand, Mo dule 2 is designed for QT dynamicity analysis in the segment of the signal previously loaded.
The filtering stage includes the following blocks, (1) a second order notch filter for removing 60 Hz power-line interference (2) a baseline drift removing algorithm using median filtering [22] and (3) a bandpass filtering stage. By default, the -3 dB bandwidth of the notch filter was set to 3 Hz which implies a quality factor (Q) of 20. The median filter uses two windows sizes of 200 ms and 600 ms respectively. The first stage, using the 200 ms window, removes the P waves and the QRS complexes of each heartbeat. The second one, using the 600 ms window, eliminates the T waves. After removing the physiological waves, the resulting signal is considered the baseline wander and consequently, it is subtracted from the original ECG signal [22]. The bandpass filtering stage consists of a fourth order forward-backward Butterworth filter with cut-off frequencies of 0.05 and 50 Hz. As an alternative to the latter, a low pass high order FIR filter is also available.
The second stage of the preprocessing is the QRS complex detection. Two QRS complex detection algorithms were provided with the tool, the first one is based on thresholds on the first and second derivatives of the ECG signal [23]. The second one is based on parabolic fitting [24].
The SQA block determines the lead which is more suitable for the further analysis, either QTVI analysis (Module 1) or QT dynamicity analysis (Module 2). In order to assess the quality of the signal, six quality indexes proposed in [25] were implemented on the software:
-
The relative power in the QRS complex (pSQI)
-
The relative power in the baseline (basSQI)
-
The baseline wander check in time domain (bsSQI)
-
The relative energy in the QRS complex (eSQI)
-
The relative amplitude of high frequency noise (hfSQI)
-
The relative standard deviation (STD) of QRS complex (rsdSQI)
In a separate experimental study (unpublished data), we observed that the above indexes increase with the quality of the signal. Since the purpose is to select the lead with the highest quality, a simple majority voting algorithm is proposed as follows. First, a counter is initialized to zero. Then, the first index is computed for both leads. If the index for lead A is greater than the index of lead B the counter is increased, otherwise is decremented. The process is repeated for all indexes. The software recommends one lead or the other one based on the values of the counter (voting index, VI), see Figure 2. The case where the counter remains equal to zero corresponds to a tie in the voting process, in such case the software will recommend the lead with the highest normalized index defined as follows,
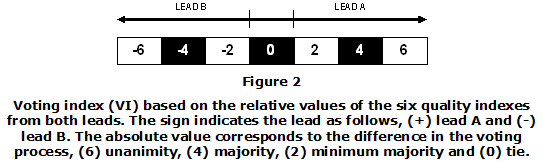
where L is the lead (A or B) and the ![]() symbol means the normalized value of the current index with respect to both leads e.g. the normalized │pSQIA│ for lead A is the ratio between pSQIA and the maximum among both pSQIA and pSQIB. The normalized pSQIB is computed in a similar way.
symbol means the normalized value of the current index with respect to both leads e.g. the normalized │pSQIA│ for lead A is the ratio between pSQIA and the maximum among both pSQIA and pSQIB. The normalized pSQIB is computed in a similar way.
In the unlikely case that both NVIA and NVIB are equal, the software would recommend using Lead A. It is important to note that the suggestion made by the software is relative to the current segments of both leads. Thus, from the point of view of the further analyses, it does not mean that the rejected lead is inappropriate nor a guarantee that the suggested one fits the quality requirements.
A key prerequisite in QT analysis is that irregular ectopic heartbeats should be avoided, i.e., the analysis should be carried out on Normal-to-Normal (NN) intervals. In the case of QTVI computation, less than the 5% of the labeled heartbeats might be ectopic. In such cases, the procedure recommends interpolating on the resampled instantaneous interval series by using a linear spline approach [6]. For QT dynamicity analysis ectopic heartbeats are not allowed. This fact points out that an irregular heartbeat detection block is needed in order to perform the analysis on NN intervals. PyECG provides two tensor-based methods for detecting irregular heartbeats. The first method uses an unsupervised approach [26]. First, a tensor (3-ways array) is constructed from the signal. Next, a rank-1 Canonical Polyadic Decomposition (CPD) takes place. Then, the mode-3 loading vector of the decomposition is used for classifying the heartbeats. Here, a threshold is constructed using the median and the standard deviation of the mode-3 loading vector. The second method uses a machine learning supervised scheme. The first stages are similar to the first method, but a Support Vector Machine (SVM) is used as the classifier. The inputs to the system are the mode-3 vectors resulting from a rank-r CPD where r > 1 [27]. Both methods can be used either separately or together. However, depending on the length of the segment under analysis it might be more efficient to use the unsupervised approach on short-term analysis. Conversely, the supervised method or the combination of both approaches might be preferable on longer segments. The combination scheme uses the output of the unsupervised classifier as preliminary inputs to the supervised method. This leads to a faster training set generation.
Since Holter systems may provide both, QRS complex detection and heartbeat labeling, it is possible to load such information directly from the recording file. Hence, the user can make use of the tools available on the software or load the corresponding annotations. Any combination of both alternatives can be used as well.
After preprocessing the signal, the tool is ready for performing the required analysis using either Module 1 or Module 2. Previously, the user is motivated to select the lead of interest. This choice should be guided by the suggestion of the software based on the signal quality assessment. However, the user can still perform the analysis on any of the two leads regardless of the software suggestion.
2.1. QTVI and qt dynamicity modules
The QTVI module performs the computation of the QTVI marker following the algorithm proposed by Berger [6]. The QTVI algorithm involves the definition of a single or several 256-second segments (epochs) and a template heartbeat in the segment under analysis. A QT interval template is generated by identifying the start of the QRS complex (Q onset, Qon) and the end of the T-wave (T offset, Toff) on the template heartbeat [6]. Given these requirements, the workflow for the QTVI module is shown in Figure 3.
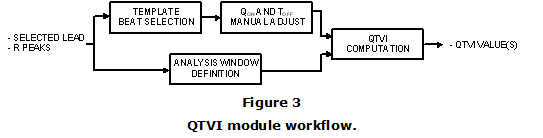
First, a template heartbeat must be manually selected by the user. Then, the software will suggest the Qon and Toff points for the current heartbeat. Both points may be manually adjusted if needed. The analysis windows can be defined by setting the starting sample at any point of the signal. This module allows computing QTVI in a single window or in a sequence of either overlapping or non-overlapping windows.
QT dynamicity analysis requires the detection of the Qon and Toff points for every heartbeat in the segment under analysis. This module includes both Qon and Toff automatic detection algorithms. The Qon detection is based on the method of the maximum triangle area. The Toff detection algorithm uses the method based on the area under the T-wave proposed by Zhang [28]. The Module 2 workflow is shown in Figure 4.
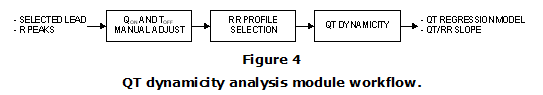
Although the software automatically detects Qon and Toff points for every heartbeat, these annotations must be manually corrected in order to guarantee the precision of the points. Hence, the first step is to check the generated points and manually correct them wherever it is needed. The next step is to select an RR profile for the QT analysis. Three profiles are available.
-
QT depends on the previous RR interval (default)
-
QT depends on a history of several previous RR intervals; this relation can be modeled by using a linear weighted (LW) average of preceding RR intervals [9].
-
Idem to the previous one but using an exponentially weighted (EW) profile [9].
The general expression for all profiles is given by the following equation,
where RRi is the RR interval for the i-th heartbeat, Niis the number of heartbeats in a time lag window of a pre-defined length and ωj are the weights of the last Ni RR intervals within the window. Different profiles can be generated by varying the weights dependence on j, e.g. a linear weighting (LW) approach is given by
As mentioned above, another feasible varying law for the weights is the exponential weighting (EW) defined as
Finally, the QT dynamicity block determines the QT/RR model and their parameters given the RR profile and the QT interval series.
3. RESULTS
Figure 5 shows the user interface of PyECG. The application includes a Main Menu and a Toolbar in the topmost side of the main screen. Both of them provide access to all the options of the software. Thus, hereinafter only the commands in the Toolbar will be commented. The Toolbar is divided in three sections. The first one includes actions related to the application management. The second and the third sections provide actions which support the general workflow previously discussed.

Section (1) provides the following actions, Open,Save, Options and Exit. The Open action (1.1) allows the user to load a Holter recording from disk, currently the tool supports Excorde E3C and Matlab® files. This command gives the alternative of loading either a segment of the signal or the whole recording. Save (1.2) allows storing the results of the analyses to a file. The button Options (1.3) provides access to configuration parameters of the preprocessing stage. Here, the user can select and configure the QRS detector and the irregular heartbeat detection algorithm. Finally, the Exit action (1.4) quits the application.
The analysis section (2) implements both processing modules. The elements on this section will be available after opening a recording. The actions included are: Detect R, Load R annotations from file, Detect irregular heartbeats, Load label annotations from file, QTVI and QT dynamicity.
The Detect R action (2.1) filters the signal and automatically detects the R peaks on both leads of the record, Figure 6. The command shows the detected points directly onto the signal plot. Two panels with possibly missing or incorrectly detected R peaks are available to the right. Both panels show the time positions where the algorithm failed and allow going directly to these positions by clicking on them. This makes easier the manual correction task, Figure 6.
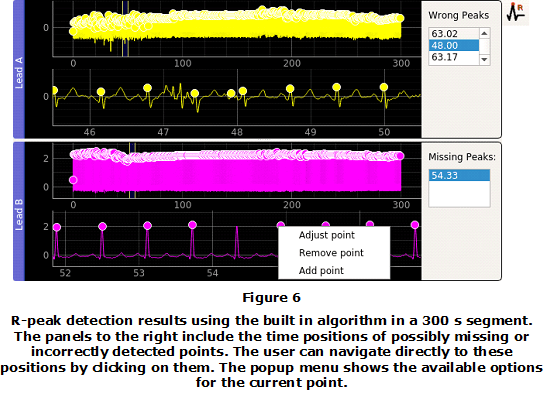
The action Load annotations from file (2.2) shows the R-peak positions of the signal, but no R-peak detection is executed by the tool. Instead, the R-peak positions are loaded from the recording file. However, still the signal is pre-filtered as was described for the Detect R action. After finishing the detection/loading process, the user can manually correct the positions in case of errors. The software provides a very simple interface for correcting the detected points. Essentially, a popup menu with three entries is available for each detected point. The possible actions include Adjust point,Remove point and Add point. Adjust point allows moving the current point around the window, Remove point deletes the current point and Add point includes a new point, Figure 6.
Together with preprocessing actions, the user can explore the power spectrum of the signal before and after filtering. Section 3 unique action, namely Spectrum (3.1) shows the power spectrum for both leads at any time it is invoked. This action is useful as reference on the effect of the filtering approach implemented in the software.
Both Detect irregular heartbeats (2.3) and Load label annotations from file (2.4) actions show the class label (regular/irregular) associated to every single heartbeat. The main goal of both actions is to provide information on the number of irregular heartbeats in the segment under analysis. Thus, the user can judge the pertinence of computing the QT indexes on this interval. The IHB is accessible via action 2.3. It processes the signal and assigns a label to each heartbeat. Depending on the algorithm a guided training process may be needed. Action 2.4 involves loading the label annotations stored on the recording file. Although the software is not intended for annotating records, it is possible to override the automatically generated labels provided by both, the built-in algorithms and the recording file.
QTVI and QT dynamicity buttons implement the Module 1 and Module 2 stages respectively. Clicking on the QTVI button will display a dock with a brief tutorial on computing QTVI. Here, the software gives details on all the necessary steps and suggests a lead for QTVI computation. Then the user should select a lead by clicking on one of the push buttons provided. Once the lead has been selected a new dock is displayed with only the lead of interest, Figure 7.
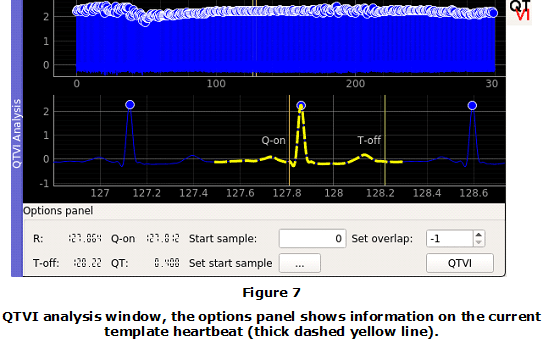
The new dock includes three sections, (1) a global view of the segment of the signal under analysis, (2) a zoomed area of the signal and (3) an information/control panel. The latter, provides the following features, (1) displays information on the current template heartbeat, (2) gives options for defining the analysis window (start and overlapping factor) and (3) allows computing QTVI, Figure 7.
The user can define the template heartbeat by clicking on any R peak displayed on the zoomed region. As response to this action, the software will change the color of the current heartbeat. Furthermore, two vertical lines will be displayed suggesting the Q-on and T-off points of the current heartbeat respectively. These lines can be manually adjusted by the user, Figure 7.
There are two alternatives for specifying the start of the 256 s analysis window. The first one is by directly writing the time in seconds or the number of the sample on the Start sample input control. The second one is a graphical alternative where the user selects the start by clicking on the push button (….) and moves the start line around the zoomed area. The Set overlap control allows for computing either one or several QTVI values on the current segment, depending on its value. A negative value corresponds to only one QTVI computation. Zero or positive values correspond to the percentages of overlapping among adjacent 256 s windows for computing each QTVI. Finally, the QTVI push button allows the user to compute the QTVI value or values given the defined parameters.
Clicking on the QT dynamicity button follows the same approach as QTVI, i.e. a brief tutorial on QT dynamicity analysis is displayed and the user is motivated to select the lead with the best relative signal quality. After selecting the lead of interest, a new three-panel dock is opened. The first panel shows the whole segment, the second one, is a zoomed version which includes automatically generated Qon and Toff points for every heartbeat on the segment. Finally, the third one is a control panel for managing options.
Every single Qon and Toff point can be manually adjusted by the user on the zoomed panel. The possible actions include, Adjust point (the point can be moved around), Add point (a new point is included, its position is given by the mean value between the current point and the previous one), Remove point (the point is deleted). All these actions are available on a contextual menu accessible by clicking on the point.
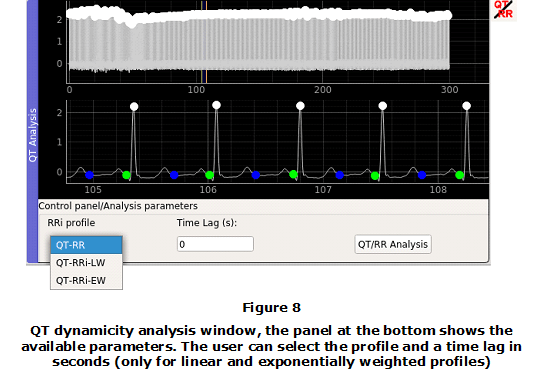
The control panel provides options for selecting one of the three available RR profiles through the combo box named RRi profile. LW and EW profiles require a time lag parameter that can be specified in the corresponding control (Time Lag), Figure 8. The QT dynamicity analysis is performed by clicking on the QT/RR button. As a response to this action, a new panel is displayed with the QT/RR analysis results. The results panel shows a QT-RR scatter plot and the fitted model. Up to 10 models of the relation QT/RR can be evaluated [9]. The parameters of each model are available on demand by clicking on the name of the model.
3.1. Tool test
This section shows a brief qualitative and quantitative evaluation of PyECG. Several tests have been performed using both, synthetic signals and recordings from the Excorde 3C Holter [29]. The tests will focus on the main features of the tool i.e. signal quality assessment, irregular heartbeats detection and QT analysis.
The SQA block was evaluated by using the tutorial feature of both modules. The tutorial explains the steps for performing the further analysis and suggests a lead for doing so. Precisely this suggestion expresses the results of the SQA module by coding the VI absolute value (VIav) in words as follows. The unanimity (VIav=6) corresponds to the words “strongly recommends”, the word “recommends” means majority (VIav=4), simple majority (VIav=2) is coded to “slightly recommends” and a tie (VIav=0) corresponds to “suggests”. The sign of VI points out the lead i.e. is positive for lead A and negative for lead B.
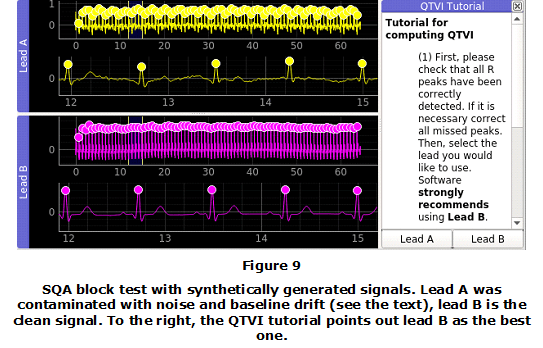
The test for SQA block evaluation consists of comparing artificial signals with different SNR. The signals were generated with the synthetic generator from McSharry et al [30]. The parameters for the model are given in Table 1. Thus, with this model a noise free signal was generated. Then, this signal was contaminated with uniformly distributed noise with amplitude 0.08 mV. Moreover, a 1 Hz, 0.3 mV baseline drift was added. Both, the contaminated and the clean signals were assembled in a single two channel signal. This two-channel record was loaded into the tool. Here, the contaminated signal corresponds to channel A and the clean one to Lead B. After detecting the R points, the SQA module can be evaluated by clicking on the QTVI action, see Figure 9. Since there is a clear difference among qualities of the channels it is expected that the software recommends Lead B, this can be seen in Figure 9.
![Table 1. McSharry et al [30] model parameters for two experiments, the SQA evaluation and the QT dynamicity analysis.](/img/revistas/eac/v39n2/t0105218.gif)
An IHB stage performance evaluation requires annotated records. Thus, eight 15-min segments of different Holter recordings were manually annotated by cardiologists using an independent tool. Table 2 shows the main characteristics of these records. The sample intends to cover several heartbeat classes such as Normal beats (NORMB), Supraventricular Ectopic Beats (SVEB), Paced Beats (PB) and Ventricular Ectopic Beats (VEB). Moreover, the recordings include signal quality changes, different types of noise, artifacts and other types of disturbances that normally affect ambulatory signals.

Table 3 shows the results of the performance evaluation using the segments of Table 2. It is worth to note that a true positive corresponds to the correct detection of an abnormal heartbeat. Excluding the positive predictive value (P+), the performance indexes are all above 96%. The worst performance index is P+ with a 64.4%. This means that the classifier is very sensitive to morphology variations. Indeed, the signal quality changes affect the morphology and consequently the output of the classifier. For instance, in segments 1 and 5 most of the false positives found, correspond to intervals where the signal quality is compromised by an increment in the noise level. We hypothesize that in such cases, better performances can be achieved by using supervised classification because the knowledge of the user can be incorporated into the classification system. However, the proper assessment of the previous hypothesis would require an extensive evaluation, which is beyond the scope of this paper. Furthermore, for the purposes of this tool, this behavior is not too critical because it is important to avoid low-quality segments. Hence, a very sensitive classifier may notify the user about poor quality segments.

In order to evaluate the performance of the detection algorithms the QT Database (QTDB) [31] was used. QTDB is publicly available in Physionet [32] and was designed for evaluating the performance of algorithms for event detection in ECG. It consists of short segments (15 min) extracted from 105 Holter recordings, each with two channels. The sampling frequency is 250 Hz.
Regarding the evaluation of the Toff detector, the following experiment was designed. First, the output of the original algorithm was obtained for all records in QTDB. This results in 65 records with at least one lead, which satisfies Group I error stratification level according to the CSE Working Party criterion [33]. This level corresponds to the records where the mean error (accuracy) on detecting Toff is below 15 ms and the standard deviation of the error (precision) is below 30.6 ms. When both leads accomplished this criterion, the best one was chosen. Then, 40 recordings were selected at random from this group for the evaluation using PyECG. The results of this evaluation are shown in Table 4.
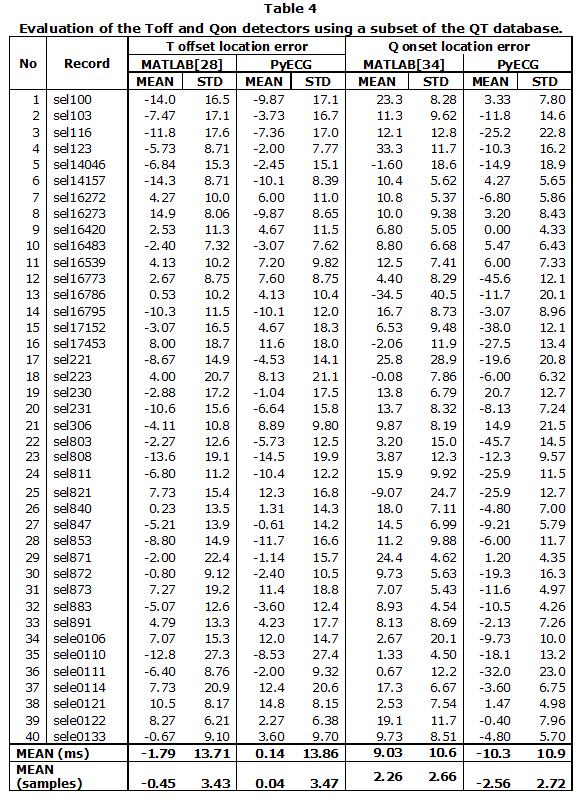
Although the original output is slightly better, both algorithms produce similar results and the overall performance barely changes. Indeed, the absolute differences in the mean accuracy and the mean precision are below one sample at 250 Hz (4 ms). Hence, it is clear that both versions have equal performances so the PyECG implementation is reliable enough and follows the original algorithm.
The Q onset detection algorithm was compared against the output of the wavelet delineator [34] included in the ECG-kit. This is a well-known delineator which is also publicly available in Phyisonet. The original code of the wavelet delineator is written in Matlab® and a similar methodology was followed for the comparison. Once more, the ground-truth is the set of Q onset annotations available on QTDB. Besides, the previous 40 records were also used and the same lead was considered for both algorithms. Although this approach for evaluating the Q onset detectors will result in lower performances, this choice is based on the fact that for the purposes of the QT/RR and QTVI analysis only one lead is finally available. Thus, all the fiducial points should be detected on the chosen lead.
From Table 4, it is noticeable that there are significant differences among algorithms. The method implemented tends to estimate the Q onset points before their real positions whereas the wavelet delineator tends to estimate the Qon points after their true location. In terms of the QT interval and assuming an exact Toff, the former method would overestimate the QT interval. Conversely, the wavelet delineator would underestimate it. Nevertheless, in terms of the absolute value of the accuracy, both methods have approximately the same mean accuracy. This is also the case of the precision parameter where the absolute difference is negligible (0.3 ms). Hence, the Q onset detector based on trapezium area has almost the same precision as the wavelet delineator; however, it is simpler and faster than the wavelet approach.
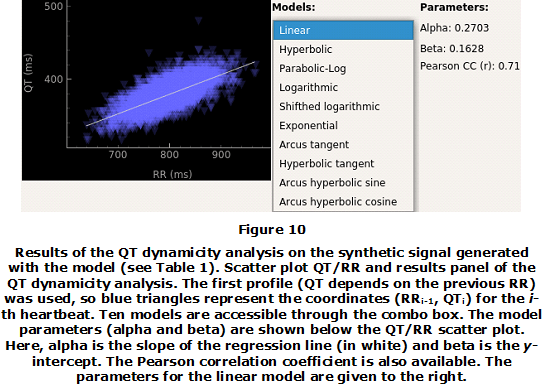
Finally, a brief test of the QT dynamicity block was performed as follows. Using the McSharry model a 3700 s signal was generated. The parameters for the model are shown in Table 1. Two signals were generated A signal with 0.01 mV of noise amplitude and a clean signal which is used as reference. This free-noise reference is used for determining the positions of the Qon and Toff points. These positions were determined outside of the tool using Matlab®. Then the QT/RR slope and y-intercept were computed using the previous points and the first profile (QT depends on the previous RR). The values of both, the slope and the y-intercept are taken as ground truth.
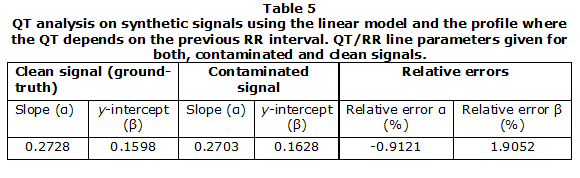
The noise-contaminated signal was loaded into the tool. The QT dynamicity analysis module was launched after filtering and R peak detection and correction processes. Since it is known that the model generates QT intervals linearly related to the RR intervals [30], the linear model for regression was used. The results for the linear model using the first profile and the relative errors with respect to the reference are given in Table 5. It is noticeable that no manual correction to the automatically generated Qon and Toff points was used. In spite of the latter, low relative errors were obtained, see Table 2. Figure 10 shows the results for this analysis using PyECG and the QT-RR scatter plot. Besides, the regression line is also visible.
So far we have described PyECG and some validation examples have been discussed. However, there is still room for improvements and the main goals for future versions of this tool will be the following:
-
To introduce a second level classifier for separating between supraventricular origin (SVEB) and ventricular origin (VEB) heartbeats in order to determine the VEB/SVEB occurrence ratio and/or to support studies on the behavior of RR or QT after a VEB
-
To provide support for several holter systems and formats, currently PyECG supports only Excorde 3C recordings and Matlab® files.
-
To include adaptive algorithms for detecting characteristic points like the end of the T wave (Toff) in order to improve the detection performance.
This tool was presented during a scientific seminar of the Cardiology Service at the Hospital Provincial Saturnino Lora, Santiago de Cuba, Cuba. We received a positive feedback from the cardiologists and the first steps for the evaluation of the tool by the service were planned. A major concern on the use of the tool was that so far PyECG runs for Linux OS and the available computers at the service run Windows. Thus, a necessary step is to port the tool to Windows OS in order to assure a fast introduction of PyECG.
4. CONCLUSIONS
This paper presented a software tool for the analysis of the QT interval on Holter recordings. The main contribution of this work is that this is the first public software tool that allows to compute indexes that have been pointed out as markers of repolarization instability. Besides, irregular heartbeats can be detected and rejected. This feature, together with a block for signal quality assessment allows for an easy, reliable and fast evaluation on the feasibility of the current segment for the QT analysis. It can be used by cardiologists and specialists with no programming skills as a tool for research on QT interval analysis. Moreover, the software has been tested with records from the Excorde 3C Holter system which is widespread throughout hospitals in Cuba.
ACKNOWLEDGMENTS
This work has been supported by the Belgian Development Cooperation through VLIR-UOS (Flemish Interuniversity Council-University Cooperation for Development) in the context of the Institutional University Cooperation programme with Universidad de Oriente. Carolina Varon is a postdoctoral fellow of the Research Foundation-Flanders (FWO). The research leading to these results has received funding from the European Research Council under the European Union's Seventh Framework Programme (FP7/2007-2013) / ERC Advanced Grant: BIOTENSORS (n° 339804). This paper reflects only the authors' views and the Union is not liable for any use that may be made of the contained information.
REFERENCES
1. Bernardo AFB, Vanderlei LCM, Garner DM, HRV Analysis: A clinical and diagnostic tool in Chronic Obstructive Pulmonary Disease. International scholarly research notices. 2014.
2. Yperzeele L, Van Hooff RJ, De Smedt A, De Keyser J, Brouns R. Heart rate variability and baroreceptor sensitivity in acute stroke: a systematic review. International Journal of Stroke. 2015;10(6):796-800.
3. Goldenberg I, Moss AJ, Zareba W, QT Interval: How to Measure It and What Is “Normal”. Journal of Cardiovascular Electrophysiology 2006:3(17)
4. Nielsen J, Fan W, Graff C, Kanters JK. QT dynamics during treatment with sertindole. Therapeutic Advances in Psychopharmacology. 2015;5(1):26-31.
5. Zareba W. Drug induced QT prolongation. Cardiology Journal, 2007;14(6):523-533.
6. Berger RD, Kasper EK, Kenneth LB, Marban E, Calkins H, Tomaseli GF. Beat-to-Beat QT Interval Variability: Novel Evidence for Repolarization Lability in Ischemic and Nonischemic Dilated Cardiomyopathy. Circulation. 1997;96:1557-1565.
7. Dobson CP, Kim A, Haigney M. QT variability index. Progress in cardiovascular diseases. 2013;56(2):186-194.
8. Sharif H, O'Leary D, Ditor D: Comparison of QT-interval and variability index methodologies in individuals with spinal cord injury. Spinal cord, 2016.
9. Pueyo E, Smetana P, Caminal P, Bayés DL A, Malik M, Laguna P, Characterization of QT interval adaptation to RR interval changes and its use as a risk-stratifier of arrhythmic mortality in amiodarone-treated survivors of acute myocardial infarction. IEEE Transactions on Biomedical Engineering. 2004;51(9):1511-1520.
10. Vázquez-Seisdedos CR, Análisis del ECG en monitorización ambulatoria para el diagnóstico cardíaco y la predicción de muerte cardíaca súbita, PhD Thesis, Universitat Politècnica de Catalunya; 2001, Available in:http://www.tdx.cat/handle/10803/6320.
11. Lederman YS, Balucani C, Lazar J, Steinberg L, Gugger J, Levine SR. Relationship between QT interval dispersion in acute stroke and stroke prognosis: a systematic review. Journal of Stroke and Cerebrovascular Diseases. 2014;23(10):2467-2478.
12. Monasterio V, Laguna P, Cygankiewicz I, Vázquez R, Bayés-Genís A, Bayés DL A, et al. Average T-wave alternans activity in ambulatory ECG records predicts sudden cardiac death in patients with chronic heart failure. Heart Rhythm. 2012;9(3):383-389.
13. Pasala T, Dettmer M, Leo PJ, Laurita KR, Kaufman ES. Microvolt T-wave alternans amplifies spatial dispersion of repolarization in human subjects with ischemic cardiomyopathy. Journal of electrocardiology. 2016;49(5):733-739.
14. Tarvainen MP, Niskanen JP, Lipponen JA, Ranta-Aho PO, Karjalainen PA. Kubios HRV--heart rate variability analysis software. Computer methods and programs in biomedicine. 2014;113(1):210-220.
15. Rodríguez-Liñares L, Lado MJ, Vila XA, Méndez AJ, Cuesta P. gHRV: heart rate variability analysis made easy. Computer methods and programs in biomedicine. 2014;116(1):26-38.
16. Rodríguez-Liñares L, Méndez AJ, Lado MJ, Olivieri DN, Vila XA, Gómez-Conde I. An open source tool for heart rate variability spectral analysis. Computer methods and programs in biomedicine. 2011;103(1):39-50.
17. Pichot V, Roche F, Celle S, Barthelemy JC, Chouchou F. HRVanalysis: a free software for analyzing cardiac autonomic activity. Frontiers in physiology. 2016;7.
18. Perakakis P, Joffily M, Taylor M, Guerra P, Vila J. KARDIA: A Matlab software for the analysis of cardiac interbeat intervals. Computer methods and programs in biomedicine. 2010;98(1):83-89.
19. Singh B, Bharti N. Software tools for heart rate variability analysis. International Journal of Recent Scientific Research. 2015;6(4):3501-3506.
20. Demski AJ, Llamedo-Soria M. ecg-kit a Matlab Toolbox for Cardiovascular Signal Processing. Journal of Open Research Software. 2016;e8, DOI: http://doi.org/10.5334/jors.86.
21. Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: Machine learning in Python. Journal of Machine Learning Research. 2011;12(Oct):2825-2830.
22. De Chazal P, Heneghan C, Sheridan E, Reilly R, Nolan P, O'Malley M. Automated processing of the single-lead electrocardiogram for the detection of obstructive sleep apnoea. IEEE Transactions on Biomedical Engineering. 2003;50(6):686-696.
23. Köhler BU, Hennig C, Orglmeister R. The Principles of Software QRS Detection: Reviewing and Comparing Algorithms for Detecting this Important ECG Waveform. IEEE Engineering in Medicine and Biology. 2002;21(1):42-57
24. Manriquez AI, Zhang Qinghua. An algorithm for QRS onset and offset detection in single lead electrocardiogram records. 2007 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society. Lyon;France;2007;541-544.
25. Li Q, Rajagopalan C, Clifford GD, A machine learning approach to multi-level ECG signal quality classification. Computer methods and programs in biomedicine. 2014;117(3):435-447.
26. Goovaerts G, De Wel O, Vandenberk B, Willems R, Van Huffel S. Detection of Irregular Heartbeats Using Tensors. Proceedings of the 42nd Annual Computing in Cardiology. Nice; France; 2015;573-576
27.Suárez-León AA, Goovaerts G, Vázquez-Seisdedos CR, Van Huffel S. ECG beat Classification Using Tensors and Support Vector Machines. Proceedings of the 43rd Annual Computing in Cardiology. Vancouver; Canada; 2016;43:1037-1040
28. Zhang Q, Manriquez AI, Médigue C, Papelier Y, Sorine M. An algorithm for robust and efficient location of T-wave ends in electrocardiograms. IEEE Transactions on Biomedical Engineering. 2006;53(12):2544-2552.
29. COMBIOMED. Excorde 3C Holter. 2009, Available in: http://www.combiomed.sld.cu/descargas/sueltos/es-/EXCORDE%203C%20Catalogo%20ESPANOL.pdf.
30. McSharry PE, Clifford GD, Lionel T, Smith LA. A Dynamical Model for Generating Synthetic Electrocardiogram Signals. IEEE Transactions on Biomedical Engineering. 2003;50(3):289-294.
31. Laguna P, Mark RG, Goldberg A, Moody GB. A database for evaluation of algorithms for measurement of QT and other waveform intervals in the ECG, in: Computers in Cardiology.1997:673-676.
32. Goldberger AL, Amaral LA, Glass L, Hausdorff JM, Ivanov PC, Mark RG, et al. Physiobank, physiotoolkit, and physionet components of a new research resource for complex physiologic signals. Circulation, 2000;101(23) e215–e220.
33. The CSE Working Party, Recommendations for measurement standards in quantitative electrocardiography, European Heart Journal. 1985; 6(10):815–825.
34. Martínez JP, Almeida R, Olmos S, Rocha AP, Laguna P. A wavelet-based ECG delineator: evaluation on standard databases, IEEE Transactions on biomedical engineering. 2004;51 (4):570–581.
Received: 30/1/2018
Approved: 25/4/2018
Alexander Alexeis Suárez-León, received the degree in automation engineering and the M.Sc. degree on biomedical engineering both from the Universidad de Oriente, Santiago de Cuba, Cuba in 2008 and 2013 respectively. His research interests include machine learning and tensor methods applied to biomedical signal processing, e-mail aasl@uo.edu.cu.














